HAEM5:T-prolymphocytic leukaemia: Difference between revisions
| [unchecked revision] | [checked revision] |
| (3 intermediate revisions by 2 users not shown) | |||
| Line 1: | Line 1: | ||
[[HAEM5:Table_of_Contents|Haematolymphoid Tumours (WHO Classification, 5th ed.)]] | [[HAEM5:Table_of_Contents|Haematolymphoid Tumours (WHO Classification, 5th ed.)]] | ||
==Primary Author(s)*== | ==Primary Author(s)*== | ||
| Line 31: | Line 29: | ||
{| class="wikitable" | {| class="wikitable" | ||
|Acceptable | |Acceptable | ||
|N/A | |N/A | ||
| Line 73: | Line 70: | ||
|Major diagnostic criteria.<ref name=":6" /> Rarely, t(X;7)(q28;q34) is observed, where the TCRβ enhancer (7q34) substitutes for TCRα/δ, leading to the same functional outcome | |Major diagnostic criteria.<ref name=":6" /> Rarely, t(X;7)(q28;q34) is observed, where the TCRβ enhancer (7q34) substitutes for TCRα/δ, leading to the same functional outcome | ||
|} | |} | ||
==Individual Region Genomic Gain/Loss/LOH== | ==Individual Region Genomic Gain/Loss/LOH== | ||
Approximately 70-80% of T-PLL karyotypes are complex, which is considered minor diagnostic criteria, and usually include 3-5 or more structural aberrations. Common cytogenetic abnormalities include those of chromosome 8, such as idic(8)(p11.2), t(8;8)(p11.2;q12), and trisomy 8q. Other frequent changes are deletions in 12p13 and 22q, gains in 8q24 (MYC), and abnormalities in chromosomes 5p, 6, and 17.<ref name=":5">Elenitoba-Johnson K, et al. T-prolymphocytic leukemia. In: WHO Classification of Tumours Editorial Board. Haematolymphoid tumours [Internet]. Lyon (France): International Agency for Research on Cancer; 2024 [cited 2024 June 12]. (WHO classification of tumors series, 5th ed.; vol. 11). Available from: https://tumourclassification.iarc.who.int/chaptercontent/63/209</ref> | Approximately 70-80% of T-PLL karyotypes are complex, which is considered minor diagnostic criteria, and usually include 3-5 or more structural aberrations. Common cytogenetic abnormalities include those of chromosome 8, such as idic(8)(p11.2), t(8;8)(p11.2;q12), and trisomy 8q. Other frequent changes are deletions in 12p13 and 22q, gains in 8q24 (MYC), and abnormalities in chromosomes 5p, 6, and 17.<ref name=":5">Elenitoba-Johnson K, et al. T-prolymphocytic leukemia. In: WHO Classification of Tumours Editorial Board. Haematolymphoid tumours [Internet]. Lyon (France): International Agency for Research on Cancer; 2024 [cited 2024 June 12]. (WHO classification of tumors series, 5th ed.; vol. 11). Available from: https://tumourclassification.iarc.who.int/chaptercontent/63/209</ref> | ||
| Line 154: | Line 149: | ||
|Minor diagnostic criteria.<ref name=":6" /> | |Minor diagnostic criteria.<ref name=":6" /> | ||
|} | |} | ||
==Diagnostic criteria== | ==Diagnostic criteria== | ||
Diagnosis requires either <u>all three major criteria</u> '''or''' the <u>first two major criteria along with one minor criterion</u>:<ref name=":5" /> | Diagnosis requires either <u>all three major criteria</u> '''or''' the <u>first two major criteria along with one minor criterion</u>:<ref name=":5" /> | ||
| Line 172: | Line 165: | ||
==Characteristic Chromosomal or Other Global Mutational Patterns== | ==Characteristic Chromosomal or Other Global Mutational Patterns== | ||
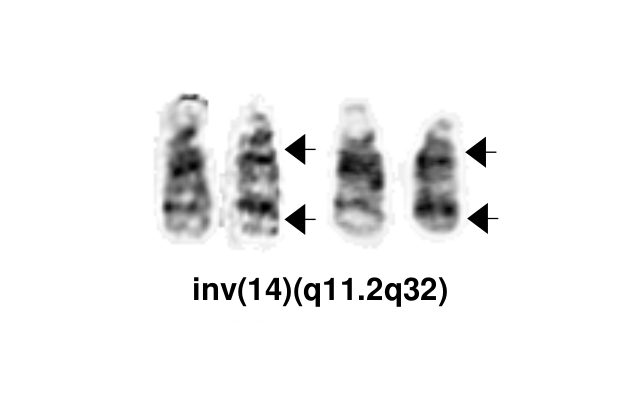
The most common chromosomal abnormality in T-PLL involves an inversion of chromosome 14, with breakpoints at q11.2 and q32.1, observed in about 60-80% of patients and described as inv(14). Additionally, in 10-20% of cases, there is a translocation t(14;14)(q11.2;q32.1).<ref name=":5" /> <ref name=":7" />[[File:Inv(14)(q11.2q32).png|thumb|Inv(14)(q11.2q32)|alt=]] | The most common chromosomal abnormality in T-PLL involves an inversion of chromosome 14, with breakpoints at q11.2 and q32.1, observed in about 60-80% of patients and described as inv(14). Additionally, in 10-20% of cases, there is a translocation t(14;14)(q11.2;q32.1).<ref name=":5" /> <ref name=":7" />[[File:Inv(14)(q11.2q32).png|thumb|Inv(14)(q11.2q32)|alt=|center]] | ||
{| class="wikitable sortable" | {| class="wikitable sortable" | ||
|- | |- | ||
| Line 197: | Line 190: | ||
|No | |No | ||
|Minor diagnostic criteria, the most frequent cooperating lesion <ref name=":6" /> | |Minor diagnostic criteria, the most frequent cooperating lesion <ref name=":6" /> | ||
|} | |} | ||
==Gene Mutations (SNV/INDEL)== | ==Gene Mutations (SNV/INDEL)== | ||
Although gene mutations beyond ''TCL1'' family alterations are not yet recognized as diagnostic criteria and remain under investigation for T-PLL, the mutational landscape of T-PLL provides valuable insights. These discoveries open up potential avenues for novel targeted therapies in treating this aggressive form of leukemia. | Although gene mutations beyond ''TCL1'' family alterations are not yet recognized as diagnostic criteria and remain under investigation for T-PLL, the mutational landscape of T-PLL provides valuable insights. These discoveries open up potential avenues for novel targeted therapies in treating this aggressive form of leukemia. | ||
| Line 213: | Line 206: | ||
|Mutation/deletion, loss of heterozygosity, or biallelic mutation | |Mutation/deletion, loss of heterozygosity, or biallelic mutation | ||
|Tumor Suppressor Gene | |Tumor Suppressor Gene | ||
|Common | |Common | ||
|D, P, T | |D, P, T | ||
|No | |No | ||
| Line 223: | Line 216: | ||
|''FBXW10'' | |''FBXW10'' | ||
<br /> | <br /> | ||
| | |Loss-of-function | ||
|Tumor Supressor Gene | |Tumor Supressor Gene | ||
|Common | |Common | ||
|Unknown | |Unknown | ||
|No | |No | ||
| Line 231: | Line 224: | ||
|- | |- | ||
|''IL2RG,'' ''JAK1, JAK3, STAT5B'' | |''IL2RG,'' ''JAK1, JAK3, STAT5B'' | ||
| | |Variable activating mutations | ||
|Oncogene | |Oncogene | ||
|Variable based on gene; Recurrent to Common | |Variable based on gene; Recurrent to Common | ||
|D, P, T | |D, P, T | ||
|No | |No | ||
| Line 247: | Line 240: | ||
|- | |- | ||
|''BCOR'' | |''BCOR'' | ||
| | |Loss-of-function | ||
|Tumor Supressor Gene | |Tumor Supressor Gene | ||
|Rare | |Rare | ||
| Line 255: | Line 248: | ||
|- | |- | ||
|''SAMHD1'' | |''SAMHD1'' | ||
| | |Loss-of-function | ||
|Tumor Supressor Gene | |Tumor Supressor Gene | ||
|Recurrent | |Recurrent | ||
| Line 263: | Line 256: | ||
|- | |- | ||
|''CHEK2'' | |''CHEK2'' | ||
| | |Loss-of-function | ||
|Tumor Supressor Gene | |Tumor Supressor Gene | ||
|Rare | |Rare | ||
|Unknown | |Unknown | ||
|No | |No | ||
|''CHEK2'' mutations may indicate a defective DNA damage response and aggressive disease <ref name=":3" /><ref>{{Cite journal|last=Braun|first=Till|last2=Dechow|first2=Annika|last3=Friedrich|first3=Gregor|last4=Seifert|first4=Michael|last5=Stachelscheid|first5=Johanna|last6=Herling|first6=Marco|date=2021|title=Advanced Pathogenetic Concepts in T-Cell Prolymphocytic Leukemia and Their Translational Impact|url=https://pubmed.ncbi.nlm.nih.gov/34869023|journal=Frontiers in Oncology|volume=11|pages=775363|doi=10.3389/fonc.2021.775363|issn=2234-943X|pmc=8639578|pmid=34869023}}</ref> | |''CHEK2'' mutations may indicate a defective DNA damage response and aggressive disease <ref name=":3" /><ref>{{Cite journal|last=Braun|first=Till|last2=Dechow|first2=Annika|last3=Friedrich|first3=Gregor|last4=Seifert|first4=Michael|last5=Stachelscheid|first5=Johanna|last6=Herling|first6=Marco|date=2021|title=Advanced Pathogenetic Concepts in T-Cell Prolymphocytic Leukemia and Their Translational Impact|url=https://pubmed.ncbi.nlm.nih.gov/34869023|journal=Frontiers in Oncology|volume=11|pages=775363|doi=10.3389/fonc.2021.775363|issn=2234-943X|pmc=8639578|pmid=34869023}}</ref> | ||
|- | |- | ||
|''TP53'' | |''TP53'' | ||
| | |Variable inactivating/loss of function mutations | ||
|Tumor Supressor Gene | |Tumor Supressor Gene | ||
|Rare | |Rare | ||
|P (may portend resistance to therapy) | |P (may portend resistance to therapy) | ||
|No | |No | ||
|Mutations in TP53 are less frequent than deletions.<ref name=":9" />May show overexpression of p53 in some cases.<ref name=":7" /> | |Mutations in TP53 are less frequent than deletions.<ref name=":9" />May show overexpression of p53 in some cases.<ref name=":7" /> | ||
| Line 371: | Line 364: | ||
==Links== | ==Links== | ||
See references. | |||
==References== | ==References== | ||
<references /> | <references /> | ||
==Notes== | ==Notes== | ||
<nowiki>*</nowiki>Primary authors will typically be those that initially create and complete the content of a page. | <nowiki>*</nowiki>''Citation of this Page'': Tizro P, Eno C, Kitahara S.“T-prolymphocytici leukemia”. Compendium of Cancer Genome Aberrations (CCGA), Cancer Genomics Consortium (CGC), updated 01/6/2026, <nowiki>https://ccga.io/index.php/HAEM5:T-prolymphocytic_leukaemia</nowiki>. | ||
<nowiki>*</nowiki>Primary authors will typically be those that initially create and complete the content of a page. If a subsequent user modifies the content and feels the effort put forth is of high enough significance to warrant listing in the authorship section, please contact the [[Leadership|''Associate Editor'']] or other CCGA representative. When pages have a major update, the new author will be acknowledged at the beginning of the page, and those who contributed previously will be acknowledged below as a prior author. | |||
Prior Author(s): N/A | |||
[[Category:HAEM5]] | [[Category:HAEM5]] | ||
[[Category:DISEASE]] | [[Category:DISEASE]] | ||
[[Category:Diseases T]] | [[Category:Diseases T]] | ||
Latest revision as of 23:06, 6 January 2026
Haematolymphoid Tumours (WHO Classification, 5th ed.)
Primary Author(s)*
Parastou Tizro, MD1, Celeste C. Eno2, PhD, Sumire Kitahara, MD2
1City of Hope, Duarte, CA
2Cedars-Sinai, Los Angeles, CA
WHO Classification of Disease
| Structure | Disease |
|---|---|
| Book | Haematolymphoid Tumours (5th ed.) |
| Category | T-cell and NK-cell lymphoid proliferations and lymphomas |
| Family | Mature T-cell and NK-cell neoplasms |
| Type | Mature T-cell and NK-cell leukaemias |
| Subtype(s) | T-prolymphocytic leukaemia |
Related Terminology
| Acceptable | N/A |
| Not Recommended | N/A |
Gene Rearrangements
Rearrangements involving the TCL1 (T-cell leukemia/lymphoma 1) family genes—TCL1A, MTCP1 (mature T-cell proliferation), or TCL1B (also known as TCL1/MTCP1-like 1 [TML1])—are highly specific to T-PLL and occur in more than 90% of cases. These translocations juxtapose the TRA locus with the oncogenes TCL1A or TCL1B, or in the case of t(X;14), with the MTCP1 gene.[1][2]
| Driver Gene | Fusion(s) and Common Partner Genes | Molecular Pathogenesis | Typical Chromosomal Alteration(s) | Prevalence -Common >20%, Recurrent 5-20% or Rare <5% (Disease) | Diagnostic, Prognostic, and Therapeutic Significance - D, P, T | Established Clinical Significance Per Guidelines - Yes or No (Source) | Clinical Relevance Details/Other Notes |
|---|---|---|---|---|---|---|---|
| inv(14) |
TCRα/δ and TCL1A | Pericentric inversion within chromosome 14, leading to juxtaposition of the TCRα/δ enhancer to the TCL1A locus and aberrant overexpression of TCL1A | inv(14)(q11.2q32.1)* | Common ~60% | D | No | These genetic abnormalities serve as diagnostic markers and generally indicate an aggressive disease. Major diagnostic criteria.[1] |
| t(14;14) | TCRα/δ and TCL1A | Reciprocal translocation between the two homologous chromosome 14s, leading to juxtaposition of the TCRα/δ enhancer to the TCL1A locus and aberrant overexpression of TCL1A | t(14;14)(q11.2;q32.1)* | Recurrent ~20-25% | D | No | *In most diagnostic/genetic reports (e.g., FISH or karyotype), the inversion and translocation may grouped together. Their distinction is mainly cytogenetic, not biological. |
| t(X;14) | TCRα/δ and MTCP1 | Transcriptional activation of MTCP1 via juxtaposition to TCRα/δ enhancer elements, leading to AKT pathway activation | t(X;14)(q28;q11.2) | Rare ~5% | D | No | Major diagnostic criteria.[1] Rarely, t(X;7)(q28;q34) is observed, where the TCRβ enhancer (7q34) substitutes for TCRα/δ, leading to the same functional outcome |
Individual Region Genomic Gain/Loss/LOH
Approximately 70-80% of T-PLL karyotypes are complex, which is considered minor diagnostic criteria, and usually include 3-5 or more structural aberrations. Common cytogenetic abnormalities include those of chromosome 8, such as idic(8)(p11.2), t(8;8)(p11.2;q12), and trisomy 8q. Other frequent changes are deletions in 12p13 and 22q, gains in 8q24 (MYC), and abnormalities in chromosomes 5p, 6, and 17.[3]
Table: A list of clinically significant and/or recurrent CNAs and CN-LOH with potential or strong diagnostic, prognostic and treatment implications in T-PLL are listed below.
| Chr # | Gain, Loss, Amp, LOH | Minimal Region Cytoband and/or Genomic Coordinates [Genome Build; Size] | Relevant Gene(s) | Diagnostic, Prognostic, and Therapeutic Significance - D, P, T | Established Clinical Significance Per Guidelines - Yes or No (Source) | Clinical Relevance Details/Other Notes |
|---|---|---|---|---|---|---|
| 8 | Gain | idic(8)(p11.2)
t(8;8)(p11.2;q12) trisomy 8q |
AGO2 [3], MYC | D | No | Recurrent secondary finding (70-80% of cases). Minor diagnostic criteria.[1] |
| 5 | Abnormality | 5p, 5q [4] | Unknown | D, P | No | Minor diagnostic criteria.[1] |
| 6 | Abnormality | Gain of 6p, loss of 6q [5][6] | Unknown | Unknown | No | |
| 11 | Loss | 11q21-q23.3 | ATM | D, P, T | No | Frequent, Minor diagnostic criteria.[1]PARP inhibitors may be considered as an off-label therapy (NCT03263637) |
| 12 | Loss | 12p13 | CDKN1B | D, P | No | Haploinsufficiency of the CDKN1B gene contributes to the development of T-PLL.[7]Minor diagnostic criteria.[1] |
| 13 | Loss | 13q14.3 | D | No | Minor diagnostic criteria.[1] | |
| 17 | Loss | 17p13 | TP53 [2] | P,T | No | May portend resistance to therapy |
| 22 | Loss | Monosomy 22 | BCL11B [9] | D | No | Minor diagnostic criteria.[1] |
Diagnostic criteria
Diagnosis requires either all three major criteria or the first two major criteria along with one minor criterion:[3]
- Major criteria:
- 5 x 109/L cells of T PLL phenotype in peripheral blood or bone marrow
- T cell clonality by molecular or flow cytometry methods
- Abnormalities of 14q32 or Xq28 or expression of TCL1A/B or MTC**
- Minor criteria:
- Abnormalities involving chromosome 11
- Abnormalities in chromosome 8
- Abnormalities in chromosome 5, 12, 13, 22 or complex karyotype
- Involvement of specific sites (spleen, effusions)
**Cases lacking these abnormalities may be referred to as "TCL1 family-negative T-PLL." by some investigators[9]. It is, however, recommended in WHO-HAEM5 that such cases should preferably be classified as peripheral T-cell lymphoma NOS with leukemic involvement (after exclusion of other specific leukemic T-cell entities) as there are insufficient clinicopathological and molecular data to determine the relationship of TCL1 family–negative T-PLL to T-PLL.
Characteristic Chromosomal or Other Global Mutational Patterns
The most common chromosomal abnormality in T-PLL involves an inversion of chromosome 14, with breakpoints at q11.2 and q32.1, observed in about 60-80% of patients and described as inv(14). Additionally, in 10-20% of cases, there is a translocation t(14;14)(q11.2;q32.1).[3] [2]
| Chromosomal Pattern | Molecular Pathogenesis | Prevalence -
Common >20%, Recurrent 5-20% or Rare <5% (Disease) |
Diagnostic, Prognostic, and Therapeutic Significance - D, P, T | Established Clinical Significance Per Guidelines - Yes or No (Source) | Clinical Relevance Details/Other Notes |
|---|---|---|---|---|---|
| inv(14)(q11q32)
t(14;14)(q11.2;q32.1) |
TCL1A or MTCP1 activation with constitutive activation of the AKT signaling pathway, promoting proliferation and survival | Common | D | No | Major diagnostic criteria [1] |
| ATM loss/mutation; del(11)(q22–q23) | Impaired DNA damage response → genomic instability, accumulation of secondary lesions | Common | D,P | No | Minor diagnostic criteria, the most frequent cooperating lesion [1] |
Gene Mutations (SNV/INDEL)
Although gene mutations beyond TCL1 family alterations are not yet recognized as diagnostic criteria and remain under investigation for T-PLL, the mutational landscape of T-PLL provides valuable insights. These discoveries open up potential avenues for novel targeted therapies in treating this aggressive form of leukemia.
| Gene | Genetic Alteration | Tumor Suppressor Gene, Oncogene, Other | Prevalence -
Common >20%, Recurrent 5-20% or Rare <5% (Disease) |
Diagnostic, Prognostic, and Therapeutic Significance - D, P, T | Established Clinical Significance Per Guidelines - Yes or No (Source) | Clinical Relevance Details/Other Notes |
|---|---|---|---|---|---|---|
| ATM
|
Mutation/deletion, loss of heterozygosity, or biallelic mutation | Tumor Suppressor Gene | Common | D, P, T | No | Since alterations at the ATM locus are found in up to 80% to 90% of T-PLL cases, it can serve as a minor diagnostic criterion.[1][10]
|
| FBXW10
|
Loss-of-function | Tumor Supressor Gene | Common | Unknown | No | |
| IL2RG, JAK1, JAK3, STAT5B | Variable activating mutations | Oncogene | Variable based on gene; Recurrent to Common | D, P, T | No | Targeting this pathway with specific JAK/STAT pathway inhibitors, has shown promise in preclinical studies and early clinical trials. Combining JAK/STAT inhibitors with other treatments, like BCL-2 inhibitors, may enhance therapeutic efficacy and improve outcomes for T-PLL patients [12][13] |
| EZH2 | Loss-of-function | Both oncogene and Tumor Suppressor Gene | Recurrent | Unknown | No | EZH2 inhibitors like tazemetostat have shown efficacy in other hematologic malignancies, providing a rationale for off-label use in T-PLL |
| BCOR | Loss-of-function | Tumor Supressor Gene | Rare | Unknown | No | A negative impact on overall survival (OS) was not observed for T-PLL patients in the study. However, this might be attributable to the relatively low number of cases compared to studies on AML and MDS.[14] |
| SAMHD1 | Loss-of-function | Tumor Supressor Gene | Recurrent | D, P | No | SAMHD1 mutations may indicate a defective DNA damage response and aggressive disease [15] |
| CHEK2 | Loss-of-function | Tumor Supressor Gene | Rare | Unknown | No | CHEK2 mutations may indicate a defective DNA damage response and aggressive disease [11][16] |
| TP53 | Variable inactivating/loss of function mutations | Tumor Supressor Gene | Rare | P (may portend resistance to therapy) | No | Mutations in TP53 are less frequent than deletions.[14]May show overexpression of p53 in some cases.[2] |
Note: A more extensive list of mutations can be found in cBioportal, COSMIC, and/or other databases. When applicable, gene-specific pages within the CCGA site directly link to pertinent external content.
Epigenomic Alterations
Research indicates that epigenetic modifications in the regulatory regions of key oncogenes and genes involved in DNA damage response and T-cell receptor regulation are clearly present. These changes are closely associated with the transcriptional dysregulation that forms the core lesions of T-PLL.[17]
Genes and Main Pathways Involved
The key pathways involved in the pathogenesis of T-cell prolymphocytic leukemia (T-PLL) include DNA damage repair, T-cell receptor (TCR) signaling, and epigenetic modulation. Additionally, there is frequent mutational activation of the IL2RG-JAK1-JAK3-STAT5B pathway, which plays a significant role in the disease's development and progression.[1]
| Gene; Genetic Alteration | Pathway | Pathophysiologic Outcome |
|---|---|---|
| TCL1A/B rearrangement | AKT signaling and TCR signal amplification pathways | Increased cell survival and proliferation |
| MTCP1 | AKT signaling and TCR signal amplification pathways | Increased cell survival and proliferation |
| ATM, CHEK2 | DNA damage repair pathway | Genomic instability |
| JAK1, JAK3, STAT5B | JAK-STAT pathway | Unchecked cell growth and survival |
| IL2RG | JAK-STAT pathway, Cytokine signaling | Promoting lymphocyte proliferation |
| EZH2 | Transcription regulator | Altering the epigenetic landscape |
Genetic Diagnostic Testing Methods
Diagnosing T-PLL involves a combination of clinical evaluation, laboratory tests, imaging studies, and genetic testing to identify diagnostic criteria. T-cell clonality can be confirmed through PCR, NGS, or flow cytometry.[18] Patients with T-PLL often exhibit complex karyotypes with recurrent genetic features that aid in diagnosis. Therefore, cytogenetic studies are useful for distinguishing T-PLL from other T-lymphoproliferative disorders.[1]
- Cytogenetic Analysis
- Karyotyping: To identify characteristic chromosomal abnormalities, such as inv(14)(q11q32), t(14;14)(q11;q32), or other translocations involving chromosome 14. Major diagnostic criteria
- Fluorescence In Situ Hybridization (FISH): To detect specific genetic abnormalities, such as TCL1 gene rearrangements as a Major diagnostic criterion or MYC as a Minor diagnostic criterion (alternatively, molecular studies could be used). see note.
Note: TCL1A break-apart probes specific to the 14q32 region can identify translocations involving TCL1A. When a TCL1A rearrangement is not identified and the patient has T-cell prolymphocytic leukemia/lymphoma (T-PLL), reflex testing using the TRAD break-apart probe set may be performed.
- Molecular Genetic Testing
- Polymerase Chain Reaction (PCR) and Reverse Transcription PCR (RT-PCR): To show the rearrangements of the TR gene (TCRB, TCRG loci) as a Major diagnostic criterion, and alternative to FISH rearrangements of the TCL1 or MTCP genes at the TRD locus can be detected by PCR. Major diagnostic criteria
- Next generation sequencing (NGS)-See note
Note: Although alterations of TCL1A, TCL1B (TML1), or MTCP are present in more than 90% of cases, they are not found in 100% of cases. Taken together, assessment of clonal TCR rearrangement, cytogenetics, and FISH are relevant genetic tests to establish the diagnosis of T-PLL. Genetic sequencing is currently not a diagnostic requirement; however, it may provide information regarding the underlying pathogenesis of T-PLL or might help to identify relevant prognostic subgroups.[1]
Familial Forms
While there is no noticeable familial clustering of T-cell prolymphocytic leukemia (T-PLL), a subset of cases can develop in the context of ataxia-telangiectasia (AT). AT is characterized by germline mutations in the ATM gene, and patients with AT are at an increased risk for various malignancies, including T-PLL. In these cases, biallelic inactivation of the ATM tumor suppressor gene occurs, with about 10% to 15% penetrance of the tumor phenotype by early adulthood. T-PLL represents nearly 3% of all malignancies in patients with ataxia-telangiectasia. [19] [20] [21]
Additional Information
- In T-PLL, the rapid growth of the disease necessitates immediate initiation of treatment. The most effective first-line treatment is alemtuzumab, an anti-CD52 antibody with remission rates over 80%. However, these remissions usually last only 1-2 years. To potentially extend remission, eligible patients are advised to undergo allogeneic blood stem cell transplantation (allo-SCT) during their first complete remission, which can lead to longer remission durations of over 4-5 years for 15-30% of patients. Consequently, the prognosis for T-PLL remains poor, with median overall survival times under two years and five-year survival rates below 5%. Ongoing studies are exploring molecularly targeted drugs and signaling pathway inhibitors, for routine clinical use in treating T-PLL.
This disease is defined/characterized as detailed below:
- T-prolymphocytic leukemia (T-PLL) is an aggressive form of T-cell leukemia marked by the proliferation of small to medium-sized prolymphocytes exhibiting a mature post-thymic T-cell phenotype.[3]
The epidemiology/prevalence of this disease is detailed below:
- T-PLL is an uncommon disease, accounting for approximately 2% of all mature lymphoid leukemias in adults. It mainly affects older individuals, with a median onset age of 65 years, ranging from 30 to 94 years. The disorder exhibits a slight male predominance, with a male to female ratio of 1.33:1.[3]
The clinical features of this disease are detailed below:
- The most prevalent symptom of the disease is a leukemic presentation, characterized by a rapid, exponential increase in lymphocyte counts, which exceed 100 × 10^9/L in 75% of patients. Approximately 30% of patients may initially experience an asymptomatic, slow-progressing phase, but this typically develops into an active disease state.[3][1]
Signs and symptoms - B symptoms (Fever, night sweats, weight loss); Hepatosplenomegaly (Frequently observed); Generalized lymphadenopathy with slightly enlarged lymph nodes (Frequently observed); Cutaneous involvement (20%); Malignant effusions (15%)
Laboratory findings - Anemia and thrombocytopenia; Marked lymphocytosis > 100 × 10^9/L (75% of cases); Atypical lymphocytosis > 5 × 10^9/L; Serum lactate dehydrogenase (LDH) (increased-may reflect disease burden); β2 microglobulin (B2M) (increased-may reflect disease burden)
The sites of involvement of this disease are detailed below:
- Peripheral blood, bone marrow, spleen (mostly red pulp), liver, lymph node (mostly paracortical), and sometimes skin and serosa (primarily pleura). Extra lymphatic and extramedullary atypical manifestations including skin, muscles and intestines are particularly common in relapse.[3]
The morphologic features of this disease are detailed below:
- Blood smears in T-PLL typically reveal anemia, thrombocytopenia, and leukocytosis, with atypical lymphocytes in three morphological forms: The most common form (75% of cases) features medium-sized cells with a high nuclear-to-cytoplasmic ratio, moderately condensed chromatin, a single visible nucleolus, and slightly basophilic cytoplasm. In 20% of cases, the cells appear as a small cell variant with densely condensed chromatin and an inconspicuous nucleolus. About 5% of cases exhibit a cerebriform variant with irregular nuclei resembling those in mycosis fungoides. Regardless of the nuclear features, a common morphological characteristic is the presence of cytoplasmic protrusions or blebs.[22]Bone marrow aspirates show clusters of these neoplastic cells, with a mixed pattern of involvement including diffuse and interstitial, in trephine core biopsy.[1]
The immunophenotype of this disease is detailed below:
- Cytochemistry: T-cell prolymphocytes show strong staining with alpha-naphthyl acetate esterase and acid phosphatase, presenting a distinctive dot-like pattern, but cytochemistry is not commonly used for diagnosis.[23]
- Immunophenotype: T-cell prolymphocytes exhibit a post-thymic T-cell phenotype. In 60% of cases, the cells are CD4+ and CD8-. In 25% of cases, they co-express both CD4 and CD8, while the remaining 15% are CD4- and CD8+.[2]
Positive (universal) - cyTCL1 (highest specificity), CD2, CD3 (may be weak), CD5, CD7 (strong), TCR-α/β, S100 (30% of cases)
Positive (subset) - CD4 (in some cases CD4+/CD8+ or CD4-/CD8+), CD52 (usually expressed at high density, therapeutic target), activation markers are variable (CD25, CD38, CD43, CD26, CD27)
Negative (universal) - TdT, CD1a, CD57, CD16, HTLV1
Negative (subset) - CD8 (in some cases CD4+/CD8+ or CD4-/CD8+)
Links
See references.
References
- ↑ 1.00 1.01 1.02 1.03 1.04 1.05 1.06 1.07 1.08 1.09 1.10 1.11 1.12 1.13 1.14 1.15 1.16 Staber, Philipp B.; et al. (2019-10-03). "Consensus criteria for diagnosis, staging, and treatment response assessment of T-cell prolymphocytic leukemia". Blood. 134 (14): 1132–1143. doi:10.1182/blood.2019000402. ISSN 1528-0020. PMC 7042666 Check
|pmc=value (help). PMID 31292114. - ↑ 2.0 2.1 2.2 2.3 2.4 Matutes E, et al., (2017). T-cell prolymphocytic leukemia, in World Health Organization Classification of Tumours of Haematopoietic and Lymphoid Tissues, Revised 4th edition. Swerdlow SH, Campo E, Harris NL, Jaffe ES, Pileri SA, Stein H, Thiele J, Arber DA, Hasserjian RP, Le Beau MM, Orazi A, and Siebert R, Editors. Revised 4th Edition. IARC Press: Lyon, France, p346-347.
- ↑ 3.0 3.1 3.2 3.3 3.4 3.5 3.6 3.7 Elenitoba-Johnson K, et al. T-prolymphocytic leukemia. In: WHO Classification of Tumours Editorial Board. Haematolymphoid tumours [Internet]. Lyon (France): International Agency for Research on Cancer; 2024 [cited 2024 June 12]. (WHO classification of tumors series, 5th ed.; vol. 11). Available from: https://tumourclassification.iarc.who.int/chaptercontent/63/209
- ↑ Tirado, Carlos A.; et al. (2012-08-20). ""T-cell prolymphocytic leukemia (T-PLL), a heterogeneous disease exemplified by two cases and the important role of cytogenetics: a multidisciplinary approach"". Experimental Hematology & Oncology. 1 (1): 21. doi:10.1186/2162-3619-1-21. ISSN 2162-3619. PMC 3514161. PMID 23211026.
- ↑ Dearden, Claire (2012-07-19). "How I treat prolymphocytic leukemia". Blood. 120 (3): 538–551. doi:10.1182/blood-2012-01-380139. ISSN 1528-0020. PMID 22649104.
- ↑ Soulier, J.; et al. (2001-07). "A complex pattern of recurrent chromosomal losses and gains in T-cell prolymphocytic leukemia". Genes, Chromosomes & Cancer. 31 (3): 248–254. doi:10.1002/gcc.1141. ISSN 1045-2257. PMID 11391795. Check date values in:
|date=(help) - ↑ Le Toriellec, Emilie; et al. (2008-02-15). "Haploinsufficiency of CDKN1B contributes to leukemogenesis in T-cell prolymphocytic leukemia". Blood. 111 (4): 2321–2328. doi:10.1182/blood-2007-06-095570. ISSN 0006-4971. PMID 18073348.
- ↑ Stengel, Anna; et al. (2014-12-06). "A Comprehensive Cytogenetic and Molecular Genetic Characterization of Patients with T-PLL Revealed Two Distinct Genetic Subgroups and JAK3 Mutations As an Important Prognostic Marker". Blood. 124 (21): 1639–1639. doi:10.1182/blood.v124.21.1639.1639. ISSN 0006-4971.
- ↑ 9.0 9.1 9.2 Fang, Hong; et al. (2023-09). "T-prolymphocytic leukemia: TCL1 or MTCP1 rearrangement is not mandatory to establish diagnosis". Leukemia. 37 (9): 1919–1921. doi:10.1038/s41375-023-01956-3. ISSN 1476-5551. PMID 37443196 Check
|pmid=value (help). Check date values in:|date=(help) - ↑ 10.0 10.1 Schrader, A.; et al. (2018-02-15). "Actionable perturbations of damage responses by TCL1/ATM and epigenetic lesions form the basis of T-PLL". Nature Communications. 9 (1): 697. doi:10.1038/s41467-017-02688-6. ISSN 2041-1723. PMC 5814445. PMID 29449575.
- ↑ 11.0 11.1 Kiel, Mark J.; et al. (2014-08-28). "Integrated genomic sequencing reveals mutational landscape of T-cell prolymphocytic leukemia". Blood. 124 (9): 1460–1472. doi:10.1182/blood-2014-03-559542. ISSN 1528-0020. PMC 4148768. PMID 24825865.
- ↑ Gomez-Arteaga, Alexandra; et al. (2019-07). "Combined use of tofacitinib (pan-JAK inhibitor) and ruxolitinib (a JAK1/2 inhibitor) for refractory T-cell prolymphocytic leukemia (T-PLL) with a JAK3 mutation". Leukemia & Lymphoma. 60 (7): 1626–1631. doi:10.1080/10428194.2019.1594220. ISSN 1029-2403. PMC 8162842 Check
|pmc=value (help). PMID 30997845. Check date values in:|date=(help) - ↑ . doi:10.1182/blood.v126.23.5486.5486 https://ashpublications.org/blood/article/126/23/5486/134544/Refractory-TCell-Prolymphocytic-Leukemia-with-JAK3. Missing or empty
|title=(help) - ↑ 14.0 14.1 Stengel, Anna; et al. (2016-01). "Genetic characterization of T-PLL reveals two major biologic subgroups and JAK3 mutations as prognostic marker". Genes, Chromosomes & Cancer. 55 (1): 82–94. doi:10.1002/gcc.22313. ISSN 1098-2264. PMID 26493028. Check date values in:
|date=(help) - ↑ Johansson, Patricia; et al. (2018-01-19). "SAMHD1 is recurrently mutated in T-cell prolymphocytic leukemia". Blood Cancer Journal. 8 (1): 11. doi:10.1038/s41408-017-0036-5. ISSN 2044-5385. PMC 5802577. PMID 29352181.
- ↑ Braun, Till; et al. (2021). "Advanced Pathogenetic Concepts in T-Cell Prolymphocytic Leukemia and Their Translational Impact". Frontiers in Oncology. 11: 775363. doi:10.3389/fonc.2021.775363. ISSN 2234-943X. PMC 8639578 Check
|pmc=value (help). PMID 34869023 Check|pmid=value (help). - ↑ Tian, Shulan; et al. (2021-04-15). "Epigenetic alteration contributes to the transcriptional reprogramming in T-cell prolymphocytic leukemia". Scientific Reports. 11 (1): 8318. doi:10.1038/s41598-021-87890-9. ISSN 2045-2322. PMC 8050249 Check
|pmc=value (help). PMID 33859327 Check|pmid=value (help). - ↑ Kotrova, Michaela; et al. (2018-11). "Next-generation amplicon TRB locus sequencing can overcome limitations of flow-cytometric Vβ expression analysis and confirms clonality in all T-cell prolymphocytic leukemia cases". Cytometry. Part A: The Journal of the International Society for Analytical Cytology. 93 (11): 1118–1124. doi:10.1002/cyto.a.23604. ISSN 1552-4930. PMID 30414304. Check date values in:
|date=(help) - ↑ Suarez, Felipe; et al. (2015-01-10). "Incidence, presentation, and prognosis of malignancies in ataxia-telangiectasia: a report from the French national registry of primary immune deficiencies". Journal of Clinical Oncology: Official Journal of the American Society of Clinical Oncology. 33 (2): 202–208. doi:10.1200/JCO.2014.56.5101. ISSN 1527-7755. PMID 25488969.
- ↑ Taylor, A. M.; et al. (1996-01-15). "Leukemia and lymphoma in ataxia telangiectasia". Blood. 87 (2): 423–438. ISSN 0006-4971. PMID 8555463.
- ↑ Li, Geling; et al. (2017-12-26). "T-cell prolymphocytic leukemia in an adolescent with ataxia-telangiectasia: novel approach with a JAK3 inhibitor (tofacitinib)". Blood Advances. 1 (27): 2724–2728. doi:10.1182/bloodadvances.2017010470. ISSN 2473-9529. PMC 5745136. PMID 29296924.
- ↑ Gutierrez, Marc; et al. (2023-07-28). "T-Cell Prolymphocytic Leukemia: Diagnosis, Pathogenesis, and Treatment". International Journal of Molecular Sciences. 24 (15): 12106. doi:10.3390/ijms241512106. ISSN 1422-0067. PMC PMC10419310 Check
|pmc=value (help). PMID 37569479 Check|pmid=value (help).CS1 maint: PMC format (link) - ↑ Yang, K.; et al. (1982-08). "Acid phosphatase and alpha-naphthyl acetate esterase in neoplastic and non-neoplastic lymphocytes. A statistical analysis". American Journal of Clinical Pathology. 78 (2): 141–149. doi:10.1093/ajcp/78.2.141. ISSN 0002-9173. PMID 6179423. Check date values in:
|date=(help)
Notes
*Citation of this Page: Tizro P, Eno C, Kitahara S.“T-prolymphocytici leukemia”. Compendium of Cancer Genome Aberrations (CCGA), Cancer Genomics Consortium (CGC), updated 01/6/2026, https://ccga.io/index.php/HAEM5:T-prolymphocytic_leukaemia.
*Primary authors will typically be those that initially create and complete the content of a page. If a subsequent user modifies the content and feels the effort put forth is of high enough significance to warrant listing in the authorship section, please contact the Associate Editor or other CCGA representative. When pages have a major update, the new author will be acknowledged at the beginning of the page, and those who contributed previously will be acknowledged below as a prior author.
Prior Author(s): N/A