|
|
| (22 intermediate revisions by 2 users not shown) |
| Line 1: |
Line 1: |
| {{DISPLAYTITLE:Acute myeloid leukaemia with DEK::NUP214 fusion}} | | {{DISPLAYTITLE:Acute myeloid leukaemia with DEK::NUP214 fusion}} |
| [[HAEM5:Table_of_Contents|Haematolymphoid Tumours (WHO Classification, 5th ed.)]] | | [[HAEM5:Table_of_Contents|Haematolymphoid Tumours (WHO Classification, 5th ed.)]] |
|
| |
| {{Under Construction}}
| |
|
| |
| <blockquote class='blockedit'>{{Box-round|title=Content Update To WHO 5th Edition Classification Is In Process; Content Below is Based on WHO 4th Edition Classification|This page was converted to the new template on 2023-12-07. The original page can be found at [[HAEM4:Acute Myeloid Leukemia (AML) with t(6;9)(p23;q34.1); DEK-NUP214]].
| |
| }}</blockquote>
| |
|
| |
| <span style="color:#0070C0">(General Instructions – The focus of these pages is the clinically significant genetic alterations in each disease type. This is based on up-to-date knowledge from multiple resources such as PubMed and the WHO classification books. The CCGA is meant to be a supplemental resource to the WHO classification books; the CCGA captures in a continually updated wiki-stye manner the current genetics/genomics knowledge of each disease, which evolves more rapidly than books can be revised and published. If the same disease is described in multiple WHO classification books, the genetics-related information for that disease will be consolidated into a single main page that has this template (other pages would only contain a link to this main page). Use [https://www.genenames.org/ <u>HUGO-approved gene names and symbols</u>] (italicized when appropriate), [https://varnomen.hgvs.org/ <u>HGVS-based nomenclature for variants</u>], as well as generic names of drugs and testing platforms or assays if applicable. Please complete tables whenever possible and do not delete them (add N/A if not applicable in the table and delete the examples); to add (or move) a row or column in a table, click nearby within the table and select the > symbol that appears. Please do not delete or alter the section headings. The use of bullet points alongside short blocks of text rather than only large paragraphs is encouraged. Additional instructions below in italicized blue text should not be included in the final page content. Please also see </span><u>[[Author_Instructions]]</u><span style="color:#0070C0"> and [[Frequently Asked Questions (FAQs)|<u>FAQs</u>]] as well as contact your [[Leadership|<u>Associate Editor</u>]] or [mailto:CCGA@cancergenomics.org <u>Technical Support</u>].)</span>
| |
|
| |
|
| ==Primary Author(s)*== | | ==Primary Author(s)*== |
|
| |
|
| Jennelle C. Hodge, PhD, FACMG | | Jennelle C. Hodge, PhD, FACMG |
|
| |
| __TOC__
| |
|
| |
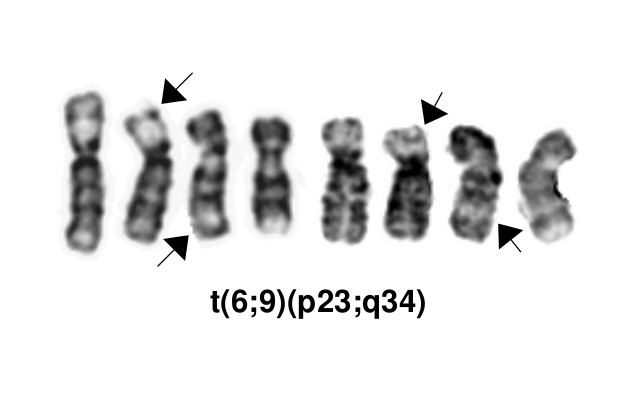
| [[File:T(6;9)(p23;q34).png]]
| |
|
| |
|
| ==WHO Classification of Disease== | | ==WHO Classification of Disease== |
| Line 39: |
Line 28: |
| |} | | |} |
|
| |
|
| ==Definition / Description of Disease== | | ==Related Terminology== |
| | |
| This is a distinct entity in the World Health Organization (WHO) classification system, and the most common associated French-American-British (FAB) classifications are M2, M4 and M1<ref name=":0">Arber DA, et al., (2017). Acute myeloid leukaemia with recurrent genetic abnormalities, in World Health Organization Classification of Tumours of Haematopoietic and Lymphoid Tissues, Revised 4th edition. Swerdlow SH, Campo E, Harris NL, Jaffe ES, Pileri SA, Stein H, Thiele J, Arber DA, Hasserjian RP, Le Beau MM, Orazi A, and Siebert R, Editors. IARC Press: Lyon, France, p137-138.</ref><ref>{{Cite journal|last=Oyarzo|first=Mauricio P.|last2=Lin|first2=Pei|last3=Glassman|first3=Armand|last4=Bueso-Ramos|first4=Carlos E.|last5=Luthra|first5=Rajyalakshmi|last6=Medeiros|first6=L. Jeffrey|date=2004|title=Acute myeloid leukemia with t(6;9)(p23;q34) is associated with dysplasia and a high frequency of flt3 gene mutations|url=https://www.ncbi.nlm.nih.gov/pubmed/15362364|journal=American Journal of Clinical Pathology|volume=122|issue=3|pages=348–358|doi=10.1309/5DGB-59KQ-A527-PD47|issn=0002-9173|pmid=15362364}}</ref>.
| |
| | |
| ==Synonyms / Terminology==
| |
| | |
| None
| |
| | |
| ==Epidemiology / Prevalence==
| |
|
| |
|
| Accounts for 0.7-1.8% of AML, occurring in both children (median age of 13 years) and adults (median age of 35-44 years)<ref name=":0" />.
| |
|
| |
| ==Clinical Features==
| |
|
| |
| Put your text here and fill in the table <span style="color:#0070C0">(''Instruction: Can include references in the table. Do not delete table.'') </span>
| |
| {| class="wikitable"
| |
| |'''Signs and Symptoms'''
| |
| |<span class="blue-text">EXAMPLE:</span> Asymptomatic (incidental finding on complete blood counts)
| |
|
| |
| <span class="blue-text">EXAMPLE:</span> B-symptoms (weight loss, fever, night sweats)
| |
|
| |
| <span class="blue-text">EXAMPLE:</span> Fatigue
| |
|
| |
| <span class="blue-text">EXAMPLE:</span> Lymphadenopathy (uncommon)
| |
| |-
| |
| |'''Laboratory Findings'''
| |
| |<span class="blue-text">EXAMPLE:</span> Cytopenias
| |
|
| |
| <span class="blue-text">EXAMPLE:</span> Lymphocytosis (low level)
| |
| |}
| |
|
| |
|
| |
| <blockquote class='blockedit'>{{Box-round|title=v4:Clinical Features|The content below was from the old template. Please incorporate above.}}</blockquote>
| |
|
| |
| Usually presents with anemia and thrombocytopenia and often with pancytopenia. In adults, the median white blood cell count is 12x10^9/L, which is generally lower than other AML types<ref name=":0" />.
| |
|
| |
| <blockquote class="blockedit">
| |
| <center><span style="color:Maroon">'''End of V4 Section'''</span>
| |
| ----
| |
| </blockquote>
| |
| ==Sites of Involvement==
| |
|
| |
| Bone marrow
| |
|
| |
| ==Morphologic Features==
| |
|
| |
| This AML subtype may present with or without monocytic features and is frequently associated with basophilia (44-62% of cases) and multilineage dysplasia (most commonly granulocytic and erythroid, and less often megakaryocytic dysplasia)<ref name=":0" />. Auer rods occur in ~1/3 of cases and ringed sideroblasts may also occur<ref name=":0" />. The peripheral blood or bone marrow have >20% blasts characterized by a non-specific myeloid immunophenotype<ref name=":0" />.
| |
|
| |
| ==Immunophenotype==
| |
|
| |
| The characteristic immunophenotype of the blasts associated with this entity is listed in the table below. In addition, basophils may present as separate clusters of CD123+, CD33+ CD38+ and HLA-DR- cells<ref name=":0" />.
| |
|
| |
| {| class="wikitable sortable"
| |
| |-
| |
| !Finding!!Marker
| |
| |-
| |
| |Positive (universal)||Myeloperoxidase (MPO), CD9, CD13, CD33, CD38 CD123, and HLA-DR
| |
| |-
| |
| |Positive (subset)||KIT(CD117), CD34, CD15, CD64 (monocyte-associated marker), and Dexoynucleotidyl Transferase (TdT)
| |
| |-
| |
| |Negative (universal)||
| |
| |-
| |
| |Negative (subset)||
| |
| |}
| |
|
| |
| ==WHO Essential and Desirable Genetic Diagnostic Criteria==
| |
| <span style="color:#0070C0">(''Instructions: The table will have the diagnostic criteria from the WHO book <u>autocompleted</u>; remove any <u>non</u>-genetics related criteria. If applicable, add text about other classification'' ''systems that define this entity and specify how the genetics-related criteria differ.'')</span>
| |
| {| class="wikitable"
| |
| |+
| |
| |WHO Essential Criteria (Genetics)*
| |
| |
| |
| |-
| |
| |WHO Desirable Criteria (Genetics)*
| |
| |
| |
| |-
| |
| |Other Classification
| |
| |
| |
| |}
| |
| <nowiki>*</nowiki>Note: These are only the genetic/genomic criteria. Additional diagnostic criteria can be found in the [https://tumourclassification.iarc.who.int/home <u>WHO Classification of Tumours</u>].
| |
| ==Related Terminology==
| |
| <span style="color:#0070C0">(''Instructions: The table will have the related terminology from the WHO <u>autocompleted</u>.)''</span>
| |
| {| class="wikitable" | | {| class="wikitable" |
| |+ | | |+ |
| |Acceptable | | |Acceptable |
| | | | |Acute myeloid leukaemia with t(6;9)(p22.3;q34.1) |
| |- | | |- |
| |Not Recommended | | |Not Recommended |
| | | | |N/A |
| |} | | |} |
|
| |
|
| ==Gene Rearrangements== | | ==Gene Rearrangements== |
| | [[File:T(6;9)(p23;q34).png]] |
|
| |
|
|
| |
| Put your text here and fill in the table <span style="color:#0070C0">(''Instructions: Details on clinical significance such as prognosis and other important information can be provided in the notes section. Please include references throughout the table. Do not delete the table.'')</span>
| |
| {| class="wikitable sortable" | | {| class="wikitable sortable" |
| |- | | |- |
| Line 141: |
Line 50: |
| !Clinical Relevance Details/Other Notes | | !Clinical Relevance Details/Other Notes |
| |- | | |- |
| |<span class="blue-text">EXAMPLE:</span> ''ABL1''||<span class="blue-text">EXAMPLE:</span> ''BCR::ABL1''||<span class="blue-text">EXAMPLE:</span> The pathogenic derivative is the der(22) resulting in fusion of 5’ BCR and 3’ABL1.||<span class="blue-text">EXAMPLE:</span> t(9;22)(q34;q11.2) | | |''DEK'' |
| |<span class="blue-text">EXAMPLE:</span> Common (CML)
| | |''DEK::NUP214''||The pathogenic derivative is the der(6) resulting in fusion of the proto-oncogene 5’ ''DEK'' and 3’''NUP214''(''CAN'').||t(6;9)(p23;q34.1) |
| |<span class="blue-text">EXAMPLE:</span> D, P, T
| | |Rare (AML) |
| |<span class="blue-text">EXAMPLE:</span> Yes (WHO, NCCN)
| | |D, P |
| |<span class="blue-text">EXAMPLE:</span>
| | |Yes (WHO) |
| The t(9;22) is diagnostic of CML in the appropriate morphology and clinical context (add reference). This fusion is responsive to targeted therapy such as Imatinib (Gleevec) (add reference). BCR::ABL1 is generally favorable in CML (add reference).
| |
| |-
| |
| |<span class="blue-text">EXAMPLE:</span> ''CIC''
| |
| |<span class="blue-text">EXAMPLE:</span> ''CIC::DUX4''
| |
| |<span class="blue-text">EXAMPLE:</span> Typically, the last exon of ''CIC'' is fused to ''DUX4''. The fusion breakpoint in ''CIC'' is usually intra-exonic and removes an inhibitory sequence, upregulating ''PEA3'' genes downstream of ''CIC'' including ''ETV1'', ''ETV4'', and ''ETV5''.
| |
| |<span class="blue-text">EXAMPLE:</span> t(4;19)(q25;q13)
| |
| |<span class="blue-text">EXAMPLE:</span> Common (CIC-rearranged sarcoma)
| |
| |<span class="blue-text">EXAMPLE:</span> D
| |
| | | |
| |<span class="blue-text">EXAMPLE:</span> | |
| | |
| ''DUX4'' has many homologous genes; an alternate translocation in a minority of cases is t(10;19), but this is usually indistinguishable from t(4;19) by short-read sequencing (add references).
| |
| |-
| |
| |<span class="blue-text">EXAMPLE:</span> ''ALK''
| |
| |<span class="blue-text">EXAMPLE:</span> ''ELM4::ALK''
| |
| | |
| | |
| Other fusion partners include ''KIF5B, NPM1, STRN, TFG, TPM3, CLTC, KLC1''
| |
| |<span class="blue-text">EXAMPLE:</span> Fusions result in constitutive activation of the ''ALK'' tyrosine kinase. The most common ''ALK'' fusion is ''EML4::ALK'', with breakpoints in intron 19 of ''ALK''. At the transcript level, a variable (5’) partner gene is fused to 3’ ''ALK'' at exon 20. Rarely, ''ALK'' fusions contain exon 19 due to breakpoints in intron 18.
| |
| |<span class="blue-text">EXAMPLE:</span> N/A
| |
| |<span class="blue-text">EXAMPLE:</span> Rare (Lung adenocarcinoma) | |
| |<span class="blue-text">EXAMPLE:</span> T | |
| |
| |
| |<span class="blue-text">EXAMPLE:</span>
| |
| | |
| Both balanced and unbalanced forms are observed by FISH (add references).
| |
| |-
| |
| |<span class="blue-text">EXAMPLE:</span> ''ABL1''
| |
| |<span class="blue-text">EXAMPLE:</span> N/A
| |
| |<span class="blue-text">EXAMPLE:</span> Intragenic deletion of exons 2–7 in ''EGFR'' removes the ligand-binding domain, resulting in a constitutively active tyrosine kinase with downstream activation of multiple oncogenic pathways.
| |
| |<span class="blue-text">EXAMPLE:</span> N/A | |
| |<span class="blue-text">EXAMPLE:</span> Recurrent (IDH-wildtype Glioblastoma)
| |
| |<span class="blue-text">EXAMPLE:</span> D, P, T
| |
| |
| |
| |
| |
| |-
| |
| | | | | |
| |
| | *This AML subtype is classified based on the presence of a t(6;9)(p23;q34.1), which results in fusion of the 5’ portion of ''DEK'' at “6p23” (specifically 6p22.3[hg38]) and the 3’ portion of ''NUP214''(''CAN'') at “9q34.1” (specifically 9q34.13[hg38])<ref name=":0">WHO Classification of Tumours Editorial Board, eds, WHO Classification of Tumours, Haematolymphoid Tumours, 5th edition, IARC Press:Lyon, 2024. Online at: [https://tumourclassification.iarc.who.int/welcome/ WHO Classification of Tumours].</ref><ref>{{Cite journal|last=Khoury|first=Joseph D.|last2=Solary|first2=Eric|last3=Abla|first3=Oussama|last4=Akkari|first4=Yassmine|last5=Alaggio|first5=Rita|last6=Apperley|first6=Jane F.|last7=Bejar|first7=Rafael|last8=Berti|first8=Emilio|last9=Busque|first9=Lambert|date=2022-07|title=The 5th edition of the World Health Organization Classification of Haematolymphoid Tumours: Myeloid and Histiocytic/Dendritic Neoplasms|url=https://pubmed.ncbi.nlm.nih.gov/35732831|journal=Leukemia|volume=36|issue=7|pages=1703–1719|doi=10.1038/s41375-022-01613-1|issn=1476-5551|pmc=9252913|pmid=35732831}}</ref>. The breakpoints are intronic, producing an in-frame fusion<ref>{{Cite journal|last=von Lindern|first=M.|last2=Fornerod|first2=M.|last3=van Baal|first3=S.|last4=Jaegle|first4=M.|last5=de Wit|first5=T.|last6=Buijs|first6=A.|last7=Grosveld|first7=G.|date=1992|title=The translocation (6;9), associated with a specific subtype of acute myeloid leukemia, results in the fusion of two genes, dek and can, and the expression of a chimeric, leukemia-specific dek-can mRNA|url=https://www.ncbi.nlm.nih.gov/pubmed/1549122|journal=Molecular and Cellular Biology|volume=12|issue=4|pages=1687–1697|doi=10.1128/mcb.12.4.1687|issn=0270-7306|pmc=PMC369612|pmid=1549122}}</ref>. The ''DEK''-''NUP214'' fusion present on the derivative chromosome 6 is considered the pathogenic entity as the reciprocal ''NUP214''-''DEK'' fusion on chromosome 9 does not appear to be transcribed<ref>{{Cite journal|last=von Lindern|first=M.|last2=Fornerod|first2=M.|last3=Soekarman|first3=N.|last4=van Baal|first4=S.|last5=Jaegle|first5=M.|last6=Hagemeijer|first6=A.|last7=Bootsma|first7=D.|last8=Grosveld|first8=G.|date=1992|title=Translocation t(6;9) in acute non-lymphocytic leukaemia results in the formation of a DEK-CAN fusion gene|url=https://www.ncbi.nlm.nih.gov/pubmed/1308167|journal=Bailliere's Clinical Haematology|volume=5|issue=4|pages=857–879|doi=10.1016/s0950-3536(11)80049-1|issn=0950-3536|pmid=1308167}}</ref>. |
| |
| | *Typically, the ''DEK''-''NUP214'' fusion presents as the sole abnormality but can be part of a complex karyotype<ref name=":0" />. |
| |
| | *Cases with the 6;9 translocation and <20% blasts are not currently classified as AML, which is controversial. Such cases should have close follow-up to monitor for development of more definitive evidence of AML or may be treated as AML if clinically appropriate<ref name=":0" />. |
| |
| | *The t(6;9) occurs in 0.6-1.7% of AML cases in children<ref name=":1" /> (REFERENCES) and about 1% of adult AML cases (REFERENCES). |
| |
| | *''DEK''::''NUP214'' has traditionally been associated with a poor prognosis in both adult and pediatric AML cases<ref name=":0" />. Of note, a 2014 retrospective analysis suggests a better outcome for pediatric patients with this translocation than previously reported<ref name=":1">{{Cite journal|last=Sandahl|first=Julie Damgaard|last2=Coenen|first2=Eva A.|last3=Forestier|first3=Erik|last4=Harbott|first4=Jochen|last5=Johansson|first5=Bertil|last6=Kerndrup|first6=Gitte|last7=Adachi|first7=Souichi|last8=Auvrignon|first8=Anne|last9=Beverloo|first9=H. Berna|date=2014|title=t(6;9)(p22;q34)/DEK-NUP214-rearranged pediatric myeloid leukemia: an international study of 62 patients|url=https://www.ncbi.nlm.nih.gov/pubmed/24441146|journal=Haematologica|volume=99|issue=5|pages=865–872|doi=10.3324/haematol.2013.098517|issn=1592-8721|pmc=4008104|pmid=24441146}}</ref>. Elevated white blood cell counts and higher bone marrow blast percentages are associated with shorter periods of overall survival and disease-free survival, respectively<ref name=":0" />. Limited data suggests early allogeneic stem cell transplantation may be associated with better overall survival compared to patients without transplantation, suggesting accurate diagnosis for these patients is crucial<ref name=":0" /><ref>{{Cite journal|last=Slovak|first=M. L.|last2=Gundacker|first2=H.|last3=Bloomfield|first3=C. D.|last4=Dewald|first4=G.|last5=Appelbaum|first5=F. R.|last6=Larson|first6=R. A.|last7=Tallman|first7=M. S.|last8=Bennett|first8=J. M.|last9=Stirewalt|first9=D. L.|date=2006|title=A retrospective study of 69 patients with t(6;9)(p23;q34) AML emphasizes the need for a prospective, multicenter initiative for rare 'poor prognosis' myeloid malignancies|url=https://www.ncbi.nlm.nih.gov/pubmed/16628187|journal=Leukemia|volume=20|issue=7|pages=1295–1297|doi=10.1038/sj.leu.2404233|issn=0887-6924|pmid=16628187}}</ref><ref>{{Cite journal|last=Ishiyama|first=K.|last2=Takami|first2=A.|last3=Kanda|first3=Y.|last4=Nakao|first4=S.|last5=Hidaka|first5=M.|last6=Maeda|first6=T.|last7=Naoe|first7=T.|last8=Taniguchi|first8=S.|last9=Kawa|first9=K.|date=2012|title=Allogeneic hematopoietic stem cell transplantation for acute myeloid leukemia with t(6;9)(p23;q34) dramatically improves the patient prognosis: a matched-pair analysis|url=https://www.ncbi.nlm.nih.gov/pubmed/21869835|journal=Leukemia|volume=26|issue=3|pages=461–464|doi=10.1038/leu.2011.229|issn=1476-5551|pmid=21869835}}</ref>. |
| |
| | *The concurrent presence of FLT3-ITD does not appear to negatively impact survival in the pediatric population<ref name=":0" />. |
| |
| |
| |}
| |
| | |
| <blockquote class='blockedit'>{{Box-round|title=v4:Chromosomal Rearrangements (Gene Fusions)|The content below was from the old template. Please incorporate above.}}</blockquote>
| |
| | |
| This AML subtype is classified based on the presence of a t(6;9)(p23;q34.1), which results in fusion of the 5’ portion of ''DEK'' at “6p23” (specifically 6p22.3[hg38]) and the 3’ portion of ''NUP214''(''CAN'') at “9q34.1” (specifically 9q34.13[hg38]). The breakpoints are intronic, producing an in-frame fusion<ref>{{Cite journal|last=von Lindern|first=M.|last2=Fornerod|first2=M.|last3=van Baal|first3=S.|last4=Jaegle|first4=M.|last5=de Wit|first5=T.|last6=Buijs|first6=A.|last7=Grosveld|first7=G.|date=1992|title=The translocation (6;9), associated with a specific subtype of acute myeloid leukemia, results in the fusion of two genes, dek and can, and the expression of a chimeric, leukemia-specific dek-can mRNA|url=https://www.ncbi.nlm.nih.gov/pubmed/1549122|journal=Molecular and Cellular Biology|volume=12|issue=4|pages=1687–1697|doi=10.1128/mcb.12.4.1687|issn=0270-7306|pmc=PMC369612|pmid=1549122}}</ref>. The ''DEK''-''NUP214'' fusion present on the derivative chromosome 6 is considered the pathogenic entity as the reciprocal ''NUP214''-''DEK'' fusion on chromosome 9 does not appear to be transcribed<ref>{{Cite journal|last=von Lindern|first=M.|last2=Fornerod|first2=M.|last3=Soekarman|first3=N.|last4=van Baal|first4=S.|last5=Jaegle|first5=M.|last6=Hagemeijer|first6=A.|last7=Bootsma|first7=D.|last8=Grosveld|first8=G.|date=1992|title=Translocation t(6;9) in acute non-lymphocytic leukaemia results in the formation of a DEK-CAN fusion gene|url=https://www.ncbi.nlm.nih.gov/pubmed/1308167|journal=Bailliere's Clinical Haematology|volume=5|issue=4|pages=857–879|doi=10.1016/s0950-3536(11)80049-1|issn=0950-3536|pmid=1308167}}</ref>. Typically the ''DEK''-''NUP214'' fusion presents as the sole abnormality but can be part of a complex karyotype<ref name=":0" />. | |
| | |
| {| class="wikitable sortable"
| |
| |-
| |
| !Chromosomal Rearrangement!!Genes in Fusion (5’ or 3’ Segments)!!Pathogenic Derivative!!Prevalence
| |
| |- | |
| |t(6;9)(p23;q34.1)||5'DEK / 3'NUP214(CAN)||der(6)||0.7-1.8% of AML | |
| |-
| |
| |} | | |} |
|
| |
| <blockquote class="blockedit">
| |
| <center><span style="color:Maroon">'''End of V4 Section'''</span>
| |
| ----
| |
| </blockquote>
| |
|
| |
|
| |
| <blockquote class='blockedit'>{{Box-round|title=v4:Clinical Significance (Diagnosis, Prognosis and Therapeutic Implications).|Please incorporate this section into the relevant tables found in:
| |
| * Chromosomal Rearrangements (Gene Fusions)
| |
| * Individual Region Genomic Gain/Loss/LOH
| |
| * Characteristic Chromosomal Patterns
| |
| * Gene Mutations (SNV/INDEL)}}</blockquote>
| |
|
| |
| This translocation has traditionally been associated with a poor prognosis in both adult and pediatric cases<ref name=":0" />. Of note, a 2014 retrospective analysis suggests a better outcome for pediatric patients with this translocation than previously reported<ref>{{Cite journal|last=Sandahl|first=Julie Damgaard|last2=Coenen|first2=Eva A.|last3=Forestier|first3=Erik|last4=Harbott|first4=Jochen|last5=Johansson|first5=Bertil|last6=Kerndrup|first6=Gitte|last7=Adachi|first7=Souichi|last8=Auvrignon|first8=Anne|last9=Beverloo|first9=H. Berna|date=2014|title=t(6;9)(p22;q34)/DEK-NUP214-rearranged pediatric myeloid leukemia: an international study of 62 patients|url=https://www.ncbi.nlm.nih.gov/pubmed/24441146|journal=Haematologica|volume=99|issue=5|pages=865–872|doi=10.3324/haematol.2013.098517|issn=1592-8721|pmc=4008104|pmid=24441146}}</ref>. Elevated white blood cell counts and higher bone marrow blast percentages are associated with shorter periods of overall survival and disease-free survival, respectively<ref name=":0" />.
| |
|
| |
| Limited data suggests early allogeneic stem cell transplantation may be associated with better overall survival compared to patients without transplantation, suggesting accurate diagnosis for these patients is crucial<ref name=":0" /><ref>{{Cite journal|last=Slovak|first=M. L.|last2=Gundacker|first2=H.|last3=Bloomfield|first3=C. D.|last4=Dewald|first4=G.|last5=Appelbaum|first5=F. R.|last6=Larson|first6=R. A.|last7=Tallman|first7=M. S.|last8=Bennett|first8=J. M.|last9=Stirewalt|first9=D. L.|date=2006|title=A retrospective study of 69 patients with t(6;9)(p23;q34) AML emphasizes the need for a prospective, multicenter initiative for rare 'poor prognosis' myeloid malignancies|url=https://www.ncbi.nlm.nih.gov/pubmed/16628187|journal=Leukemia|volume=20|issue=7|pages=1295–1297|doi=10.1038/sj.leu.2404233|issn=0887-6924|pmid=16628187}}</ref><ref>{{Cite journal|last=Ishiyama|first=K.|last2=Takami|first2=A.|last3=Kanda|first3=Y.|last4=Nakao|first4=S.|last5=Hidaka|first5=M.|last6=Maeda|first6=T.|last7=Naoe|first7=T.|last8=Taniguchi|first8=S.|last9=Kawa|first9=K.|date=2012|title=Allogeneic hematopoietic stem cell transplantation for acute myeloid leukemia with t(6;9)(p23;q34) dramatically improves the patient prognosis: a matched-pair analysis|url=https://www.ncbi.nlm.nih.gov/pubmed/21869835|journal=Leukemia|volume=26|issue=3|pages=461–464|doi=10.1038/leu.2011.229|issn=1476-5551|pmid=21869835}}</ref>.
| |
|
| |
| The concurrent presence of FLT3-ITD does not appear to negatively impact survival in the pediatric population<ref name=":0" />.
| |
|
| |
| Cases with the 6;9 translocation and <20% blasts are not currently classified as AML, which is controversial. Such cases should have close follow-up to monitor for development of more definitive evidence of AML or may be treated as AML if clinically appropriate<ref name=":0" />.
| |
|
| |
| <blockquote class="blockedit">
| |
| <center><span style="color:Maroon">'''End of V4 Section'''</span>
| |
| ----
| |
| </blockquote>
| |
| ==Individual Region Genomic Gain/Loss/LOH== | | ==Individual Region Genomic Gain/Loss/LOH== |
|
| |
|
|
| |
| Put your text here and fill in the table <span style="color:#0070C0">(''Instructions: Includes aberrations not involving gene rearrangements. Details on clinical significance such as prognosis and other important information can be provided in the notes section. Can refer to CGC workgroup tables as linked on the homepage if applicable. Please include references throughout the table. Do not delete the table.'') </span>
| |
| {| class="wikitable sortable" | | {| class="wikitable sortable" |
| |- | | |- |
| !Chr #!!'''Gain, Loss, Amp, LOH'''!!'''Minimal Region Cytoband and/or Genomic Coordinates [Genome Build; Size]'''!!'''Relevant Gene(s)''' | | !Chr #!!Gain, Loss, Amp, LOH!!Minimal Region Cytoband and/or Genomic Coordinates [Genome Build; Size]!!Relevant Gene(s) |
| !'''Diagnostic, Prognostic, and Therapeutic Significance - D, P, T''' | | !Diagnostic, Prognostic, and Therapeutic Significance - D, P, T |
| !'''Established Clinical Significance Per Guidelines - Yes or No (Source)''' | | !Established Clinical Significance Per Guidelines - Yes or No (Source) |
| !'''Clinical Relevance Details/Other Notes''' | | !Clinical Relevance Details/Other Notes |
| |-
| |
| |<span class="blue-text">EXAMPLE:</span>
| |
| 7
| |
| |<span class="blue-text">EXAMPLE:</span> Loss
| |
| |<span class="blue-text">EXAMPLE:</span>
| |
| chr7
| |
| |<span class="blue-text">EXAMPLE:</span>
| |
| Unknown
| |
| |<span class="blue-text">EXAMPLE:</span> D, P
| |
| |<span class="blue-text">EXAMPLE:</span> No
| |
| |<span class="blue-text">EXAMPLE:</span>
| |
| Presence of monosomy 7 (or 7q deletion) is sufficient for a diagnosis of AML with MDS-related changes when there is ≥20% blasts and no prior therapy (add reference). Monosomy 7/7q deletion is associated with a poor prognosis in AML (add references).
| |
| |- | | |- |
| |<span class="blue-text">EXAMPLE:</span> | | |N/A |
| 8
| |
| |<span class="blue-text">EXAMPLE:</span> Gain
| |
| |<span class="blue-text">EXAMPLE:</span>
| |
| chr8
| |
| |<span class="blue-text">EXAMPLE:</span>
| |
| Unknown
| |
| |<span class="blue-text">EXAMPLE:</span> D, P
| |
| |
| |
| |<span class="blue-text">EXAMPLE:</span>
| |
| Common recurrent secondary finding for t(8;21) (add references).
| |
| |-
| |
| |<span class="blue-text">EXAMPLE:</span>
| |
| 17
| |
| |<span class="blue-text">EXAMPLE:</span> Amp
| |
| |<span class="blue-text">EXAMPLE:</span>
| |
| 17q12; chr17:39,700,064-39,728,658 [hg38; 28.6 kb]
| |
| |<span class="blue-text">EXAMPLE:</span>
| |
| ''ERBB2''
| |
| |<span class="blue-text">EXAMPLE:</span> D, P, T
| |
| |
| |
| |<span class="blue-text">EXAMPLE:</span>
| |
| Amplification of ''ERBB2'' is associated with HER2 overexpression in HER2 positive breast cancer (add references). Add criteria for how amplification is defined.
| |
| |-
| |
| |
| |
| | | | | |
| | | | | |
| Line 283: |
Line 80: |
| | | | | |
| |} | | |} |
|
| |
| <blockquote class='blockedit'>{{Box-round|title=v4:Genomic Gain/Loss/LOH|The content below was from the old template. Please incorporate above.}}</blockquote>
| |
|
| |
| Not applicable
| |
|
| |
| <blockquote class="blockedit">
| |
| <center><span style="color:Maroon">'''End of V4 Section'''</span>
| |
| ----
| |
| </blockquote>
| |
| ==Characteristic Chromosomal or Other Global Mutational Patterns== | | ==Characteristic Chromosomal or Other Global Mutational Patterns== |
|
| |
|
|
| |
| Put your text here and fill in the table <span style="color:#0070C0">(I''nstructions: Included in this category are alterations such as hyperdiploid; gain of odd number chromosomes including typically chromosome 1, 3, 5, 7, 11, and 17; co-deletion of 1p and 19q; complex karyotypes without characteristic genetic findings; chromothripsis; microsatellite instability; homologous recombination deficiency; mutational signature pattern; etc. Details on clinical significance such as prognosis and other important information can be provided in the notes section. Please include references throughout the table. Do not delete the table.'')</span>
| |
| {| class="wikitable sortable" | | {| class="wikitable sortable" |
| |- | | |- |
| !Chromosomal Pattern | | !Chromosomal Pattern |
| !Molecular Pathogenesis | | !Molecular Pathogenesis |
| !'''Prevalence -''' | | !Prevalence - |
| '''Common >20%, Recurrent 5-20% or Rare <5% (Disease)'''
| | Common >20%, Recurrent 5-20% or Rare <5% (Disease) |
| !'''Diagnostic, Prognostic, and Therapeutic Significance - D, P, T''' | | !Diagnostic, Prognostic, and Therapeutic Significance - D, P, T |
| !'''Established Clinical Significance Per Guidelines - Yes or No (Source)''' | | !Established Clinical Significance Per Guidelines - Yes or No (Source) |
| !'''Clinical Relevance Details/Other Notes''' | | !Clinical Relevance Details/Other Notes |
| |- | | |- |
| |<span class="blue-text">EXAMPLE:</span> | | |N/A |
| Co-deletion of 1p and 18q
| |
| |<span class="blue-text">EXAMPLE:</span> See chromosomal rearrangements table as this pattern is due to an unbalanced derivative translocation associated with oligodendroglioma (add reference).
| |
| |<span class="blue-text">EXAMPLE:</span> Common (Oligodendroglioma)
| |
| |<span class="blue-text">EXAMPLE:</span> D, P
| |
| |
| |
| |
| |
| |-
| |
| |<span class="blue-text">EXAMPLE:</span>
| |
| Microsatellite instability - hypermutated
| |
| |
| |
| |<span class="blue-text">EXAMPLE:</span> Common (Endometrial carcinoma)
| |
| |<span class="blue-text">EXAMPLE:</span> P, T
| |
| |
| |
| |
| |
| |-
| |
| |
| |
| | | | | |
| | | | | |
| Line 329: |
Line 99: |
| | | | | |
| |} | | |} |
|
| |
| <blockquote class='blockedit'>{{Box-round|title=v4:Characteristic Chromosomal Aberrations / Patterns|The content below was from the old template. Please incorporate above.}}</blockquote>
| |
|
| |
| Not applicable
| |
|
| |
| <blockquote class="blockedit">
| |
| <center><span style="color:Maroon">'''End of V4 Section'''</span>
| |
| ----
| |
| </blockquote>
| |
| ==Gene Mutations (SNV/INDEL)== | | ==Gene Mutations (SNV/INDEL)== |
|
| |
|
| | *COSMIC does not have specific information on mutations related to this subtype of AML. |
|
| |
|
| Put your text here and fill in the table <span style="color:#0070C0">(''Instructions: This table is not meant to be an exhaustive list; please include only genes/alterations that are recurrent or common as well either disease defining and/or clinically significant. If a gene has multiple mechanisms depending on the type or site of the alteration, add multiple entries in the table. For clinical significance, denote associations with FDA-approved therapy (not an extensive list of applicable drugs) and NCCN or other national guidelines if applicable; Can also refer to CGC workgroup tables as linked on the homepage if applicable as well as any high impact papers or reviews of gene mutations in this entity. Details on clinical significance such as prognosis and other important information such as concomitant and mutually exclusive mutations can be provided in the notes section. Please include references throughout the table. Do not delete the table.'') </span>
| |
| {| class="wikitable sortable" | | {| class="wikitable sortable" |
| |- | | |- |
| !Gene!!'''Genetic Alteration'''!!'''Tumor Suppressor Gene, Oncogene, Other'''!!'''Prevalence -''' | | !Gene!!Genetic Alteration!!Tumor Suppressor Gene, Oncogene, Other!!Prevalence - |
| '''Common >20%, Recurrent 5-20% or Rare <5% (Disease)'''
| | Common >20%, Recurrent 5-20% or Rare <5% (Disease) |
| !'''Diagnostic, Prognostic, and Therapeutic Significance - D, P, T ''' | | !Diagnostic, Prognostic, and Therapeutic Significance - D, P, T |
| !'''Established Clinical Significance Per Guidelines - Yes or No (Source)''' | | !Established Clinical Significance Per Guidelines - Yes or No (Source) |
| !'''Clinical Relevance Details/Other Notes''' | | !Clinical Relevance Details/Other Notes |
| |- | | |- |
| |<span class="blue-text">EXAMPLE:</span>''EGFR'' | | |''FLT3'' |
|
| |
|
| <br /> | | <br /> |
| |<span class="blue-text">EXAMPLE:</span> Exon 18-21 activating mutations | | |ITD mutations |
| |<span class="blue-text">EXAMPLE:</span> Oncogene | | |Oncogene |
| |<span class="blue-text">EXAMPLE:</span> Common (lung cancer) | | |Common (AML with DEK::NUP214 fusion) |
| |<span class="blue-text">EXAMPLE:</span> T | | |<span class="blue-text">EXAMPLE:</span> T |
| |<span class="blue-text">EXAMPLE:</span> Yes (NCCN) | | |<span class="blue-text">EXAMPLE:</span> Yes (NCCN) |
| |<span class="blue-text">EXAMPLE:</span> Exons 18, 19, and 21 mutations are targetable for therapy. Exon 20 T790M variants cause resistance to first generation TKI therapy and are targetable by second and third generation TKIs (add references).
| |
| |-
| |
| |<span class="blue-text">EXAMPLE:</span> ''TP53''; Variable LOF mutations
| |
| <br />
| |
| |<span class="blue-text">EXAMPLE:</span> Variable LOF mutations
| |
| |<span class="blue-text">EXAMPLE:</span> Tumor Supressor Gene
| |
| |<span class="blue-text">EXAMPLE:</span> Common (breast cancer)
| |
| |<span class="blue-text">EXAMPLE:</span> P
| |
| |
| |
| |<span class="blue-text">EXAMPLE:</span> >90% are somatic; rare germline alterations associated with Li-Fraumeni syndrome (add reference). Denotes a poor prognosis in breast cancer.
| |
| |-
| |
| |<span class="blue-text">EXAMPLE:</span> ''BRAF''; Activating mutations
| |
| |<span class="blue-text">EXAMPLE:</span> Activating mutations
| |
| |<span class="blue-text">EXAMPLE:</span> Oncogene
| |
| |<span class="blue-text">EXAMPLE:</span> Common (melanoma)
| |
| |<span class="blue-text">EXAMPLE:</span> T
| |
| |
| |
| |
| |
| |-
| |
| |
| |
| |
| |
| |
| |
| |
| |
| |
| |
| |
| |
| | | | | |
| | *FLT3-ITD occurs in 69% of children and 78% of adults. |
| | *In contrast to FLT3-ITD mutations, FLT3-TKD is very uncommon. |
| | *The concurrent presence of FLT3-ITD with t(6;9) does not appear to negatively impact survival in the pediatric population<ref name=":0" />. |
| |}Note: A more extensive list of mutations can be found in [https://www.cbioportal.org/ <u>cBioportal</u>], [https://cancer.sanger.ac.uk/cosmic <u>COSMIC</u>], and/or other databases. When applicable, gene-specific pages within the CCGA site directly link to pertinent external content. | | |}Note: A more extensive list of mutations can be found in [https://www.cbioportal.org/ <u>cBioportal</u>], [https://cancer.sanger.ac.uk/cosmic <u>COSMIC</u>], and/or other databases. When applicable, gene-specific pages within the CCGA site directly link to pertinent external content. |
|
| |
| <blockquote class='blockedit'>{{Box-round|title=v4:Gene Mutations (SNV/INDEL)|The content below was from the old template. Please incorporate above.}}</blockquote>
| |
|
| |
| COSMIC does not have specific information on mutations related to this subtype of AML.
| |
|
| |
| ===Other Mutations===
| |
| {| class="wikitable sortable"
| |
| |-
| |
| !Type!!Gene/Region/Other
| |
| |-
| |
| |Concomitant Mutations||FLT3-ITD (69% of children and 78% of adults)
| |
| |-
| |
| |Secondary Mutations||
| |
| |-
| |
| |Mutually Exclusive||FLT3-TKD is very uncommon
| |
| |}
| |
|
| |
| <blockquote class="blockedit">
| |
| <center><span style="color:Maroon">'''End of V4 Section'''</span>
| |
| ----
| |
| </blockquote>
| |
| ==Epigenomic Alterations== | | ==Epigenomic Alterations== |
|
| |
|
| Line 411: |
Line 129: |
|
| |
|
| ==Genes and Main Pathways Involved== | | ==Genes and Main Pathways Involved== |
|
| |
|
| |
| Put your text here and fill in the table <span style="color:#0070C0">(''Instructions: Please include references throughout the table. Do not delete the table.)''</span>
| |
| {| class="wikitable sortable" | | {| class="wikitable sortable" |
| |- | | |- |
| !Gene; Genetic Alteration!!Pathway!!Pathophysiologic Outcome | | !Gene; Genetic Alteration!!Pathway!!Pathophysiologic Outcome |
| |- | | |- |
| |<span class="blue-text">EXAMPLE:</span> ''BRAF'' and ''MAP2K1''; Activating mutations | | |''DEK::NUP214'' |
| |<span class="blue-text">EXAMPLE:</span> MAPK signaling | | |Unknown |
| |<span class="blue-text">EXAMPLE:</span> Increased cell growth and proliferation | | |The fusion protein is known to act as an aberrant transcription factor, alter nuclear transport and induce myeloid cell-specific global protein synthesis<ref name=":0" /><ref>{{Cite journal|last=Ageberg|first=Malin|last2=Drott|first2=Kristina|last3=Olofsson|first3=Tor|last4=Gullberg|first4=Urban|last5=Lindmark|first5=Anders|date=2008|title=Identification of a novel and myeloid specific role of the leukemia-associated fusion protein DEK-NUP214 leading to increased protein synthesis|url=https://www.ncbi.nlm.nih.gov/pubmed/18181180|journal=Genes, Chromosomes & Cancer|volume=47|issue=4|pages=276–287|doi=10.1002/gcc.20531|issn=1098-2264|pmid=18181180}}</ref>. |
| |-
| |
| |<span class="blue-text">EXAMPLE:</span> ''CDKN2A''; Inactivating mutations | |
| |<span class="blue-text">EXAMPLE:</span> Cell cycle regulation | |
| |<span class="blue-text">EXAMPLE:</span> Unregulated cell division | |
| |- | |
| |<span class="blue-text">EXAMPLE:</span> ''KMT2C'' and ''ARID1A''; Inactivating mutations | |
| |<span class="blue-text">EXAMPLE:</span> Histone modification, chromatin remodeling | |
| |<span class="blue-text">EXAMPLE:</span> Abnormal gene expression program | |
| |-
| |
| |
| |
| |
| |
| |
| |
| |} | | |} |
|
| |
| <blockquote class='blockedit'>{{Box-round|title=v4:Genes and Main Pathways Involved|The content below was from the old template. Please incorporate above.}}</blockquote>
| |
|
| |
| The molecular mechanism is not completely understood, but the fusion protein is known to act as an aberrant transcription factor, alter nuclear transport and induce myeloid cell-specific global protein synthesis<ref name=":0" /><ref>{{Cite journal|last=Ageberg|first=Malin|last2=Drott|first2=Kristina|last3=Olofsson|first3=Tor|last4=Gullberg|first4=Urban|last5=Lindmark|first5=Anders|date=2008|title=Identification of a novel and myeloid specific role of the leukemia-associated fusion protein DEK-NUP214 leading to increased protein synthesis|url=https://www.ncbi.nlm.nih.gov/pubmed/18181180|journal=Genes, Chromosomes & Cancer|volume=47|issue=4|pages=276–287|doi=10.1002/gcc.20531|issn=1098-2264|pmid=18181180}}</ref>.
| |
|
| |
| <blockquote class="blockedit">
| |
| <center><span style="color:Maroon">'''End of V4 Section'''</span>
| |
| ----
| |
| </blockquote>
| |
| ==Genetic Diagnostic Testing Methods== | | ==Genetic Diagnostic Testing Methods== |
|
| |
|
| Karyotype, FISH, RT-PCR | | Karyotype, FISH, RT-PCR (and any other fusion detecting technologies) |
|
| |
|
| ==Familial Forms== | | ==Familial Forms== |
| Line 462: |
Line 156: |
|
| |
|
| ==References== | | ==References== |
| (use the "Cite" icon at the top of the page) <span style="color:#0070C0">(''Instructions: Add each reference into the text above by clicking where you want to insert the reference, selecting the “Cite” icon at the top of the wiki page, and using the “Automatic” tab option to search by PMID to select the reference to insert. If a PMID is not available, such as for a book, please use the “Cite” icon, select “Manual” and then “Basic Form”, and include the entire reference. To insert the same reference again later in the page, select the “Cite” icon and “Re-use” to find the reference; DO NOT insert the same reference twice using the “Automatic” tab as it will be treated as two separate references. The reference list in this section will be automatically generated and sorted''</span><span style="color:#0070C0">''.''</span><span style="color:#0070C0">)</span> <references />
| | <references /> |
| | |
| '''
| |
|
| |
|
| ==Notes== | | ==Notes== |
| Line 473: |
Line 165: |
| | | |
| <nowiki>*</nowiki>''Citation of this Page'': “Acute myeloid leukaemia with DEK::NUP214 fusion”. Compendium of Cancer Genome Aberrations (CCGA), Cancer Genomics Consortium (CGC), updated {{REVISIONMONTH}}/{{REVISIONDAY}}/{{REVISIONYEAR}}, <nowiki>https://ccga.io/index.php/HAEM5:Acute_myeloid_leukaemia_with_DEK::NUP214_fusion</nowiki>. | | <nowiki>*</nowiki>''Citation of this Page'': “Acute myeloid leukaemia with DEK::NUP214 fusion”. Compendium of Cancer Genome Aberrations (CCGA), Cancer Genomics Consortium (CGC), updated {{REVISIONMONTH}}/{{REVISIONDAY}}/{{REVISIONYEAR}}, <nowiki>https://ccga.io/index.php/HAEM5:Acute_myeloid_leukaemia_with_DEK::NUP214_fusion</nowiki>. |
| [[Category:HAEM5]][[Category:DISEASE]][[Category:Diseases A]] | | [[Category:HAEM5]] |
| | [[Category:DISEASE]] |
| | [[Category:Diseases A]] |