HAEM5:B-lymphoblastic leukaemia/lymphoma with BCR::ABL1-like features: Difference between revisions
| [pending revision] | [pending revision] |
Mark.Evans (talk | contribs) mNo edit summary |
Mark.Evans (talk | contribs) |
||
| (4 intermediate revisions by the same user not shown) | |||
| Line 12: | Line 12: | ||
==Primary Author(s)*== | ==Primary Author(s)*== | ||
Mark G. Evans, MD, Caris Life Sciences | Mark G. Evans, MD, Caris Life Sciences | ||
Sumire K. Kitahara, MD, Cedars-Sinai Medical Center | Sumire K. Kitahara, MD, Cedars-Sinai Medical Center | ||
| Line 64: | Line 62: | ||
|'''Comment''' | |'''Comment''' | ||
|- | |- | ||
| rowspan="12" | | | rowspan="12" |''[[ABL1]]'' | ||
(9q34) | (9q34) | ||
|''CENPC1'' | |''CENPC1'' | ||
| Line 150: | Line 148: | ||
| | | | ||
|- | |- | ||
| rowspan="3" | | | rowspan="3" |''[[ABL2]]'' | ||
(1q25.2) | (1q25.2) | ||
|''PAG1'' | |''PAG1'' | ||
| Line 173: | Line 171: | ||
| | | | ||
|- | |- | ||
| rowspan="2" | | | rowspan="2" |''[[CRLF2]]'' | ||
(Xp22.3 & Yp11.3) | (Xp22.3 & Yp11.3) | ||
|''[[IGH]]'' | |''[[IGH]]'' | ||
| Line 190: | Line 188: | ||
| | | | ||
|- | |- | ||
| rowspan="3" | | | rowspan="3" |''CSF1R'' | ||
(5q32) | (5q32) | ||
|''MEF2D'' | |''MEF2D'' | ||
| Line 213: | Line 211: | ||
| | | | ||
|- | |- | ||
| | |''DGKH'' (13q14.1) | ||
|''ZFAND3'' | |''ZFAND3'' | ||
|t(6;13)(p21.2;q14.1) | |t(6;13)(p21.2;q14.1) | ||
| Line 221: | Line 219: | ||
|Requires complex rearrangement due to incompatible orientation of genes with respect to chromosome arms | |Requires complex rearrangement due to incompatible orientation of genes with respect to chromosome arms | ||
|- | |- | ||
| rowspan="4" | | | rowspan="4" |''EPOR'' (19p13.2) | ||
|''[[IGH]]'' | |''[[IGH]]'' | ||
|ins(14;19)(q32;p13.2p13.2) | |ins(14;19)(q32;p13.2p13.2) | ||
| Line 250: | Line 248: | ||
| | | | ||
|- | |- | ||
| | |''IL2RB'' (22q12.3) | ||
|''MYH9'' | |''MYH9'' | ||
|22q12.3 rearrangement | |22q12.3 rearrangement | ||
| Line 258: | Line 256: | ||
|On the same chromosome arm; however, a simple deletion cannot cause the fusion due to the orientation of genes | |On the same chromosome arm; however, a simple deletion cannot cause the fusion due to the orientation of genes | ||
|- | |- | ||
| rowspan="22" | | | rowspan="22" |''[[JAK2]]'' | ||
(9p24.1) | (9p24.1) | ||
|''ATF7IP'' | |''ATF7IP'' | ||
| Line 414: | Line 412: | ||
| | | | ||
|- | |- | ||
| | |''[[PDGFRA]]'' | ||
(4q12) | (4q12) | ||
|''FIP1L1'' | |''FIP1L1'' | ||
| Line 423: | Line 421: | ||
|Interstitial deletion. Seen also in myeloid/lymphoid neoplasms with eosinophilia | |Interstitial deletion. Seen also in myeloid/lymphoid neoplasms with eosinophilia | ||
|- | |- | ||
| rowspan="8" | | | rowspan="8" |''[[PDGFRB]]'' (5q32) | ||
|''ATF7IP'' | |''ATF7IP'' | ||
|t(5;12)(q32;p13.1) | |t(5;12)(q32;p13.1) | ||
| Line 480: | Line 478: | ||
| | | | ||
|- | |- | ||
| rowspan="3" | | | rowspan="3" |''PTK2B'' (8p21.2) | ||
|''[[KDM6A]]'' | |''[[KDM6A]]'' | ||
|t(X;8)(p11.3;p21.2) | |t(X;8)(p11.3;p21.2) | ||
| Line 502: | Line 500: | ||
| | | | ||
|- | |- | ||
| rowspan="3" | | | rowspan="3" |''TYK2'' (19p13.2) | ||
|''MYB'' | |''MYB'' | ||
|t(6;19)(q23.3;p13.2) | |t(6;19)(q23.3;p13.2) | ||
| Line 667: | Line 665: | ||
|Unknown | |Unknown | ||
|No | |No | ||
|Half of cases with ''CRLF2'' overexpression have activating mutations in ''JAK1'' or ''JAK2'' that promote downstream JAK-STAT signaling;<ref name=":10" /> the most common mutation, p.R683G, occurs in the pseudokinase domain of ''JAK2'', and less common ''JAK1'' alterations have been detected, which include p.V658F most frequently. | |Half of cases with ''CRLF2'' overexpression have activating mutations in ''JAK1'' or ''JAK2'' that promote downstream JAK-STAT signaling;<ref name=":10" /> the most common mutation, p.R683G, occurs in the pseudokinase domain of ''JAK2'', and less common ''JAK1'' alterations have been detected, which include p.V658F most frequently; clinical trials examining the treatment effects of targeting JAK proteins are currently ongoing.<ref>{{Cite journal|last=Goulart|first=Hannah|last2=Jabbour|first2=Elias|last3=Short|first3=Nicholas J.|last4=Kadia|first4=Tapan M.|last5=Pemmaraju|first5=Naveen|last6=Takahashi|first6=Koichi|last7=Ravandi|first7=Farhad|last8=Konopleva|first8=Marina|last9=Jain|first9=Nitin|date=2025-11|title=A Phase I/II Trial of Ruxolitinib with Chemotherapy for Patients with Relapsed and/or Refractory Philadelphia-like Acute Lymphoblastic Leukemia|url=https://pubmed.ncbi.nlm.nih.gov/40500616|journal=Clinical Lymphoma, Myeloma & Leukemia|volume=25|issue=11|pages=800–807|doi=10.1016/j.clml.2025.05.013|issn=2152-2669|pmid=40500616}}</ref> | ||
|- | |- | ||
|''IL7R'' | |''IL7R'' | ||
| Line 679: | Line 677: | ||
|''SH2B3'' | |''SH2B3'' | ||
''IL2RB'' | ''IL2RB'' | ||
''TYK2'' | ''TYK2'' | ||
''TLSP'' | ''TLSP'' | ||
|Activating mutations | |Activating mutations | ||
| Line 709: | Line 709: | ||
|These result in B-cell progenitor proliferation; may be responsive to TKIs.<ref>{{Cite journal|last=Senapati|first=Jayastu|last2=Jabbour|first2=Elias|last3=Konopleva|first3=Marina|last4=Short|first4=Nicholas J.|last5=Tang|first5=Guilin|last6=Daver|first6=Naval|last7=Kebriaei|first7=Partow|last8=Kadia|first8=Tapan|last9=Pemmaraju|first9=Naveen|date=2023-05|title=Philadelphia-Like Genetic Rearrangements in Adults With B-Cell ALL: Refractoriness to Chemotherapy and Response to Tyrosine Kinase Inhibitor in ABL Class Rearrangements|url=https://pubmed.ncbi.nlm.nih.gov/37196217|journal=JCO precision oncology|volume=7|pages=e2200707|doi=10.1200/PO.22.00707|issn=2473-4284|pmc=10309573|pmid=37196217}}</ref> | |These result in B-cell progenitor proliferation; may be responsive to TKIs.<ref>{{Cite journal|last=Senapati|first=Jayastu|last2=Jabbour|first2=Elias|last3=Konopleva|first3=Marina|last4=Short|first4=Nicholas J.|last5=Tang|first5=Guilin|last6=Daver|first6=Naval|last7=Kebriaei|first7=Partow|last8=Kadia|first8=Tapan|last9=Pemmaraju|first9=Naveen|date=2023-05|title=Philadelphia-Like Genetic Rearrangements in Adults With B-Cell ALL: Refractoriness to Chemotherapy and Response to Tyrosine Kinase Inhibitor in ABL Class Rearrangements|url=https://pubmed.ncbi.nlm.nih.gov/37196217|journal=JCO precision oncology|volume=7|pages=e2200707|doi=10.1200/PO.22.00707|issn=2473-4284|pmc=10309573|pmid=37196217}}</ref> | ||
|- | |- | ||
|''CRLF2'' overexpression; mutations of ''CRLF2'', ''JAK1'', ''IL7R, SH2B3, IL2RB, TYK2,'' and ''TLSP''; ''JAK2'' and ''EPOR'' rearrangements | |''CRLF2'' overexpression; mutations of ''CRLF2'', ''JAK1/2'', ''IL7R, SH2B3, IL2RB, TYK2,'' and ''TLSP''; ''JAK2'' and ''EPOR'' rearrangements | ||
|JAK-STAT signaling | |JAK-STAT signaling | ||
|These potentiate the JAK2-signal transducer and upregulate the transcription 5 pathway;<ref name=":8" /> other mutations not in ''CRLF2'' and ''IL7R'' cause constitutive JAK/STAT activation downstream of CRLF2. | |These potentiate the JAK2-signal transducer and upregulate the transcription 5 pathway;<ref name=":8" /> other mutations not in ''CRLF2'' and ''IL7R'' cause constitutive JAK/STAT activation downstream of CRLF2. | ||
Latest revision as of 16:08, 6 November 2025
Haematolymphoid Tumours (WHO Classification, 5th ed.)
| This page is under construction |
editContent Update To WHO 5th Edition Classification Is In Process; Content Below is Based on WHO 4th Edition ClassificationThis page was converted to the new template on 2023-12-07. The original page can be found at HAEM4:B-Lymphoblastic Leukemia/Lymphoma, BCR-ABL1-Like.
(General Instructions – The focus of these pages is the clinically significant genetic alterations in each disease type. This is based on up-to-date knowledge from multiple resources such as PubMed and the WHO classification books. The CCGA is meant to be a supplemental resource to the WHO classification books; the CCGA captures in a continually updated wiki-stye manner the current genetics/genomics knowledge of each disease, which evolves more rapidly than books can be revised and published. If the same disease is described in multiple WHO classification books, the genetics-related information for that disease will be consolidated into a single main page that has this template (other pages would only contain a link to this main page). Use HUGO-approved gene names and symbols (italicized when appropriate), HGVS-based nomenclature for variants, as well as generic names of drugs and testing platforms or assays if applicable. Please complete tables whenever possible and do not delete them (add N/A if not applicable in the table and delete the examples); to add (or move) a row or column in a table, click nearby within the table and select the > symbol that appears. Please do not delete or alter the section headings. The use of bullet points alongside short blocks of text rather than only large paragraphs is encouraged. Additional instructions below in italicized blue text should not be included in the final page content. Please also see Author_Instructions and FAQs as well as contact your Associate Editor or Technical Support.)
Primary Author(s)*
Mark G. Evans, MD, Caris Life Sciences
Sumire K. Kitahara, MD, Cedars-Sinai Medical Center
WHO Classification of Disease
| Structure | Disease |
|---|---|
| Book | Haematolymphoid Tumours (5th ed.) |
| Category | B-cell lymphoid proliferations and lymphomas |
| Family | Precursor B-cell neoplasms |
| Type | B-lymphoblastic leukaemias/lymphomas |
| Subtype(s) | B-lymphoblastic leukaemia/lymphoma with BCR::ABL1-like features |
Related Terminology
| Acceptable | Philadelphia-like (Ph-like) B-ALL; BCR::ABL1-like B-ALL/LBL |
| Not Recommended | N/A |
Gene Rearrangements
B-lymphoblastic leukaemia/lymphoma with BCR::ABL1-like features traditionally required diagnosis by gene expression (GEX) profiling[1][2] and was found to exhibit a GEX profile similar to Philadelphia chromosome-positive B-lymphoblastic leukaemia/lymphoma but lacking BCR::ABL1. The WHO[3] and ICC[4] have since recognized recurring genomic alterations associated with B-lymphoblastic leukaemia/lymphoma with BCR::ABL1-like features, including ABL-class rearrangements, JAK-STAT activating alterations, and others. Proper identification of this disease is important, as patients may respond to targeted therapies like tyrosine kinase inhibitors (TKIs);[5] however, as most reports feature only single cases and limited series, consensus on the diagnostic/prognostic/therapeutic significance of the various genomic alterations has not been reached and currently being established.
Table derived from Akkari et al., 2020 [6] with permission from Cancer Genetics summarizes the important gene rearrangements associated with B-lymphoblastic leukaemia/lymphoma with BCR::ABL1-like features.
| 3’ Partner | 5’ Partner | Chromosome rearrangement | Gene fusion | Visible by G-banding | References | Comment |
| ABL1
(9q34) |
CENPC1 | t(4;9)(q13;q34) | CENPC1::ABL1 | YES | [7] | Requires complex rearrangement due to incompatible orientation of genes with respect to chromosome arms |
| ETV6 | t(9;12)(q34;p13) | ETV6::ABL1 | NO | [8] | Requires complex rearrangement due to incompatible orientation of genes with respect to chromosome arms | |
| FOXP1 | t(3;9)(p13;q34) | FOXP1::ABL1 on der(3) | YES | [9] | ||
| LSM14A | t(9;19)(q34;q13.1) | LSM14A::ABL1 on der(19) | YES | [7] | ||
| NUP153 | t(6;9)(p22.3;q34) | NUP153::ABL1 on der(6) | YES | [7] | ||
| NUP214 | dup(9)(q34.1q34.1) | NUP214::ABL1 | NO | [10] | Tandem duplication (~370 kb) detectable by CMA | |
| RANBP2 | t(2;9)(q12.3;q34) | RANBP::ABL1 on der(2) | YES | [5] | ||
| RCSD1 | t(1;9)(q24.2;q34) | RCSD1::ABL1 on der(1) | YES | [11] | ||
| SFPQ | t(1;9)(p34.3;q34) | SFPQ::ABL1 on der(1) | YES | [12] | ||
| SNX1 | t(9;15)(q34;q22.3) | SNX1::ABL1 on der(15) | YES | [13] | ||
| SNX2 | t(5;9)(q23.2;q34) | SNX2::ABL1 on der(5) | YES | [14] | ||
| ZMIZ1 | t(9;10)(q34;q22.3) | ZMIZ1::ABL1 on der(10) | YES | [15] | ||
| ABL2
(1q25.2) |
PAG1 | t(1;8)(q25.2;q21.1) | PAG1::ABL2 on der(1) | YES | [5] | |
| RCSD1 | 1q24.2q25.2 rearrangement | RCSD1::ABL2 | NO | [16] | On the same chromosome arm; however, a simple deletion cannot cause the fusion due to the orientation of genes | |
| ZC3HAV1 | t(1;7)(q25.2;q34) | ZC3HAV1::ABL2 on der(1) | YES | [17] | ||
| CRLF2
(Xp22.3 & Yp11.3) |
IGH | t(X;14)(p22.3;q32) or
t(Y;14)(p11.3;q32) |
IGH::CRLF2 | NO | [18][5] | |
| P2RY8 | del(X)(p22.3p22.3) or del(Y)(p11.3p11.3) | P2RY8::CRLF2 | NO | [18][5] | ||
| CSF1R
(5q32) |
MEF2D | t(1;5)(q22;q32) | MEF2D::CSF1R on der(5) | YES | [19] | |
| SSBP2 | 5q14.1q32 rearrangement | SSBP2::CSF1R | YES | [7] | On the same chromosome arm; however, a simple deletion cannot cause the fusion due to the orientation of genes | |
| TBL1XR1 | t(3;5)(q26.3;q32) | TBL1XR1::CSF1R on der(5) | YES | [7] | ||
| DGKH (13q14.1) | ZFAND3 | t(6;13)(p21.2;q14.1) | ZFAND3::DGKH | YES | [5] | Requires complex rearrangement due to incompatible orientation of genes with respect to chromosome arms |
| EPOR (19p13.2) | IGH | ins(14;19)(q32;p13.2p13.2) | IGH/EPOR | Cryptic insertion | [20] | |
| IGK | ins(2;19)(p11.2;p13.2p13.2) | IGK/EPOR | Cryptic insertion | [20] | ||
| LAIR1 | inv(19)(p13.2q13.42) | LAIR1::EPOR | NO | [20] | Inversion of chromosome 19 juxtaposes EPOR to the upstream region of LAIR1 | |
| THADA | t(2;19)(p21;p13.2) | THADA::EPOR | YES | [13] | ||
| IL2RB (22q12.3) | MYH9 | 22q12.3 rearrangement | MYH9::IL2RB | NO | [5] | On the same chromosome arm; however, a simple deletion cannot cause the fusion due to the orientation of genes |
| JAK2
(9p24.1) |
ATF7IP | t(9;12)(p24.1;p13.1) | ATF7IP::JAK2 on der(9) | NO | [5][21] | |
| BCR | t(9;22)(p24.1;q11.2) | BCR::JAK2 | ? YES | [22] | Seen also in myeloproliferative neoplasms. Requires complex rearrangement due to incompatible orientation of genes with respect to chromosome arms | |
| EBF1 | t(5;9)(q33.3;p24.1) | EBF1::JAK2 on der(9) | NO (SUBTLE) | [23] | ||
| ETV6 | t(9;12)(p24.1;p13.2) | ETV6::JAK2 on der(9) | NO (SUBTLE) | [24][25] | ||
| GOLGA5 | t(9;14)(p24.1;q32.1) | GOLGA5::JAK2 | NO (SUBTLE) | [26] | Requires complex rearrangement due to incompatible orientation of genes with respect to chromosome arms | |
| HMBOX1 | t(8;9)(p21.1;p24.1) | HMBOX1::JAK2 on der(9) | YES | [27] | ||
| OFD1 | t(X;9)(p22.2;p24.1) | OFD1::JAK2 on der(9) | NO (SUBTLE) | [28] | ||
| PAX5 | inv(9)(p13.2p24.1) | PAX5::JAK2 | YES | [29] | An inversion is required as genes are oriented in opposite directions | |
| PCM1 | t(8;9)(p22;p24.1) | PCM1::JAK2 on der(9) | YES (SUBTLE) | [13] | Seen also in myeloid/lymphoid neoplasms with eosinophilia | |
| PPFIBP1 | t(9;12)(p24.1;p11.2) | PPFIBP1::JAK2 on der(9) | YES | [13] | ||
| RFX3 | inv(9)(p24.1p24.2) | RFX3::JAK2 | NO | [7] | An inversion is required as genes are oriented in opposite directions | |
| SMU1 | inv(9)(p21.1p24.1) | SMU1::JAK2 | NO | [27] | An inversion is required as genes are oriented in opposite directions | |
| SNX29 | t(9;16)(p24.1;p13.1) | SNX29::JAK2 on der(9) | YES | [27] | ||
| SPAG9 | t(9;17)(p24.1;q21.3) | SPAG9::JAK2 on der(9) | YES | [30] | ||
| SSBP2 | t(5;9)(q14.1;p24.1) | SSBP2::JAK2 on der(9) | YES | [31] | ||
| STRN3 | t(9;14)(p24.1;q12) | STRN3::JAK2 on der(9) | YES | [32] | ||
| TERF2 | t(9;16)(p24.1;q22.1) | TERF2::JAK2 on der(9) | YES | [33] | ||
| TPR | t(1;9)(q31.1;p24.1) | TPR::JAK2 on der(9) | YES | [5] | ||
| USP25 | t(9;21)(p24.1;q21.1) | USP25::JAK2 | ? YES | [7] | Requires complex rearrangement due to incompatible orientation of genes with respect to chromosome arms | |
| ZBTB46 | t(9;20)(p24.1;q13.3) | ZBTB46::JAK2 on der(9) | NO | [13] | ||
| ZNF274 | t(9;19)(p24.1;q13.4) | ZNF274::JAK2 | NO | [7] | Requires complex rearrangement due to incompatible orientation of genes with respect to chromosome arms | |
| ZNF340 | t(9;20)(p24.1;q13.3) | ZNF340::JAK2 on der(9) | NO | [13] | ||
| PDGFRA
(4q12) |
FIP1L1 | del(4)(q12q12) | FIP1L1::PDGFRA | NO | [27] | Interstitial deletion. Seen also in myeloid/lymphoid neoplasms with eosinophilia |
| PDGFRB (5q32) | ATF7IP | t(5;12)(q32;p13.1) | ATF7IP::PDGFRB on der(5) | YES | [34][35][36] | |
| EBF1 | del(5)(q32q33.3) | EBF1::PDGFRB | NO | [37] | Interstitial deletion | |
| ETV6 | t(5;12)(q32;p13.2) | ETV6::PDGFRB on der(5) | YES | [13] | ||
| SNX29 | t(5;16)(q32;p13.1) | SNX29::PDGFRB on der(5) | YES | [13] | ||
| SSBP2 | t(5;5)(q14.1;q32) | SSBP2::PDGFRB | ? YES | [13] | On the same chromosome arm; however, a simple deletion cannot cause the fusion due to the orientation of genes | |
| TNIP1 | del(5)(q32q33.1) | TNIP1::PDGFRB | NO | [13] | Interstitial deletion. Seen also in myeloid/lymphoid neoplasms with eosinophilia | |
| ZEB2 | t(2;5)(q22.3;q32) | ZEB2::PDGFRB on der(5) | YES | [5] | ||
| ZMYND8 | t(5;20)(q32;q13.1) | ZMYND8::PDGFRB on der(5) | YES | [7] | ||
| PTK2B (8p21.2) | KDM6A | t(X;8)(p11.3;p21.2) | KDM6A::PTK2B on der(8) | YES | [5] | |
| STAG2 | t(X;8)(q25;p21.2) | STAG2::PTK2B | YES | [5] | Requires complex rearrangement due to incompatible orientation of genes with respect to chromosome arms | |
| TMEM2 | t(8;9)(p21.2;q21.1) | TMEM2::PTK2B on der(8) | YES | [13] | ||
| TYK2 (19p13.2) | MYB | t(6;19)(q23.3;p13.2) | MYB::TYK2 on der(6) | YES | [23] | |
| SMARCA4 | inv(19)(p13.2p13.2) | SMARCA4::TYK2 | NO | [13] | ||
| ZNF340 | t(19;20)(p13.2;q13.3) | ZNF340::TYK2 | NO | [13] | Requires complex rearrangement due to incompatible orientation of genes with respect to chromosome arms |
Individual Region Genomic Gain/Loss/LOH
| Chr # | Gain, Loss, Amp, LOH | Minimal Region Cytoband and/or Genomic Coordinates [Genome Build; Size] | Relevant Gene(s) | Diagnostic, Prognostic, and Therapeutic Significance - D, P, T | Established Clinical Significance Per Guidelines - Yes or No (Source) | Clinical Relevance Details/Other Notes |
|---|---|---|---|---|---|---|
| 5 | Loss | chr5:158,695,920-159,099,916
[GRCh38/hg38] |
EBF1 | Unknown | No | Deletion of EBF1 results in altered B-cell development.[38] |
| 7 | Loss | chr7:50,303,455-50,405,101
[GRCh38/hg38] |
IKZF1 | P | Yes, NCCN - Acute Lymphoblastic leukaemia | Monoallelic (often partial) deletion of the IKAROS transcription factor, encoded by IKZF1, is one of the most frequently observed genetic abnormalities in B-lymphoblastic leukaemia/lymphoma with BCR::ABL1-like features, although this finding is not specific and not included in the definition;[39] IKZF1 deletion is associated with poor prognosis.[40] |
| 9 | Loss | chr9:21,967,752-21,995,324
chr9:22,002,903-22,009,313 [GRCh38/hg38] |
CDKN2A
CDKN2B |
Unknown | No | Deletion of CDKN2A/B results in altered B-cell development.[38] |
| 9 | Loss | chr9:36,833,269-37,034,268
[GRCh38/hg38] |
PAX5 | Unknown | No | Deletion of PAX5 results in altered B-cell development.[38] |
| 12 | Loss | chr12:11,649,674-11,895,377
[GRCh38/hg38] |
ETV6 | Unknown | No | Deletion of ETV6 results in altered B-cell development.[38] |
| 13 | Loss | chr13:48,303,744-48,599,436
[GRCh38/hg38] |
RB1 | Unknown | No | Deletion of RB1 results in disrupted cell-cycle regulation.[38] |
| 17 | Loss | chr17:7,661,779-7,687,546
[GRCh38/hg38] |
TP53 | Unknown | No | Deletion of TP53 results in in disrupted cell-cycle regulation.[38] |
Characteristic Chromosomal or Other Global Mutational Patterns
| Chromosomal Pattern | Molecular Pathogenesis | Prevalence -
Common >20%, Recurrent 5-20% or Rare <5% (Disease) |
Diagnostic, Prognostic, and Therapeutic Significance - D, P, T | Established Clinical Significance Per Guidelines - Yes or No (Source) | Clinical Relevance Details/Other Notes |
|---|---|---|---|---|---|
| Chromosome X/Y cryptic deletion or translocation | These changes cause CRLF2 overexpression, upregulating the JAK-STAT pathway. | Common (>20%) | P | No | Chromosome X/Y abnormalities include either translocation of the immunoglobin heavy chain enhance locus into CRLF2 (IGH::CRLF2—more commonly seen in adults) or a cryptic deletion involving the PAR1 psuedoautosomal region, resulting in fusion of CRLF2 and P2RY8 (more commonly seen in children); these alterations involving CRLF2 have been associated with poor survival;[41] very rare alternative translocations involving CRLF2 have also been observed. |
| Polysomy or iAMP21 | These changes stem from telomere attrition that results in amplification of all or a region of chromosome 21. | Rare (<5%) | P | No | iAMP21 is considered high-risk cytogenetic abnormality/poor prognostic indicator, but it is not specific to B-lymphoblastic leukaemia/lymphoma with BCR::ABL1-like features and can be seen in other B-lymphoblastic leukaemia/lymphomas.[42] |
[Abnormal fluorescence in situ hybridization (FISH) results in interphase nuclei from a bone marrow sample using the CRLF2 dual-color, break-apart (Cytocell) and IGH dual-color, break-apart probes, reflective of IGH::CRLF2 rearrangement]
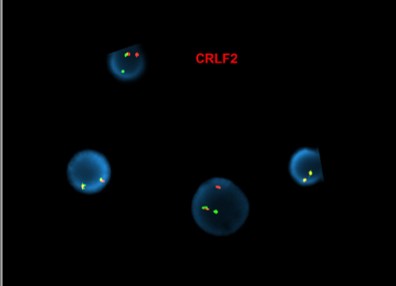
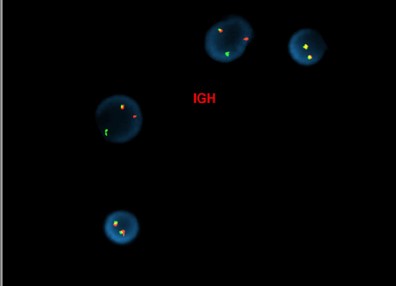
[Concurrent abnormal karyotype with trisomy 21 and a translocation involving chromosomes X, 14, and 2 in 9 of 13 cells available for analysis; metaphase FISH with the IGH break-apart probe (Vysis) confirms the presence of 5’ IGH (green signal) on the abnormal chromosome Xp33.1 (CRLF2 locus), highly suggestive on an IGH::CRLF2 fusion rearrangement: 47,XX,+21c[4]/47,idem,der(X)t(X;14)(p33.1;q32),der(2)t(2;14)(p11.2;q11.2)t(X;14),der(14)t(2;14)[5]/46,XX[4].ish der(X)(5’IGH+),der(2)(3’IGH+)]

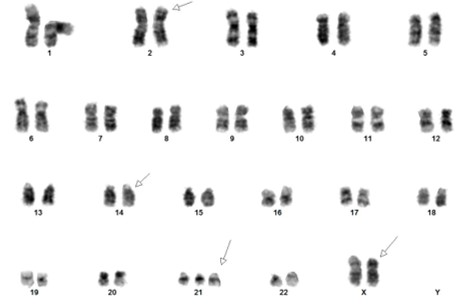
(Images courtesy of Fabiola Quintero-Rivera, MD)
Gene Mutations (SNV/INDEL)
| Gene | Genetic Alteration | Tumor Suppressor Gene, Oncogene, Other | Prevalence -
Common >20%, Recurrent 5-20% or Rare <5% (Disease) |
Diagnostic, Prognostic, and Therapeutic Significance - D, P, T | Established Clinical Significance Per Guidelines - Yes or No (Source) | Clinical Relevance Details/Other Notes |
|---|---|---|---|---|---|---|
| CRLF2 | p.F232C | Oncogene | Recurrent (5-20%) | Unknown | No | p.F232C is a gain-of-function mutation that results in constitutive dimerization and cytokine independent growth within the JAK-STAT pathway.[43] |
| JAK1
JAK2 |
p.V658Fp.R683G | Oncogene | Recurrent (5-20%) | Unknown | No | Half of cases with CRLF2 overexpression have activating mutations in JAK1 or JAK2 that promote downstream JAK-STAT signaling;[13] the most common mutation, p.R683G, occurs in the pseudokinase domain of JAK2, and less common JAK1 alterations have been detected, which include p.V658F most frequently; clinical trials examining the treatment effects of targeting JAK proteins are currently ongoing.[44] |
| IL7R | Activating mutations | Oncogene | Recurrent (5-20%) | Unknown | No | IL7R is the partner gene of CRLF2; gain-of-function mutations potentiate CRFL2 and its cofactor IL7RA forming a receptor for thymic stromal-derived lymphopoietin, leading to JAK-STAT activation.[45] |
| SH2B3
IL2RB TYK2 TLSP |
Activating mutations | Oncogene | Recurrent (5-20%) | Unknown | No | These result in constitutive activation of JAK-STAT signaling and are often present as multi-subclonal (suggestive of secondary driver events).[46] |
| RAS pathway genes | Activating mutations | Oncogenes | Recurrent (5-20%) | Unknown | No | Activating mutations in KRAS, NF1, PTPN11, and other genes upregulate the MAP kinase pathway and have been found at a higher frequency in B-lymphoblastic leukaemia/lymphoma with BCR::ABL1-like features compared to other B-lymphoblastic leukaemia/lymphomas.[47] |
Note: A more extensive list of mutations can be found in cBioportal, COSMIC, and/or other databases. When applicable, gene-specific pages within the CCGA site directly link to pertinent external content
Epigenomic Alterations
Not applicable
Genes and Main Pathways Involved
| Gene; Genetic Alteration | Pathway | Pathophysiologic Outcome |
|---|---|---|
| ABL-class rearrangements | Tyrosine kinase signaling | These result in B-cell progenitor proliferation; may be responsive to TKIs.[48] |
| CRLF2 overexpression; mutations of CRLF2, JAK1/2, IL7R, SH2B3, IL2RB, TYK2, and TLSP; JAK2 and EPOR rearrangements | JAK-STAT signaling | These potentiate the JAK2-signal transducer and upregulate the transcription 5 pathway;[45] other mutations not in CRLF2 and IL7R cause constitutive JAK/STAT activation downstream of CRLF2. |
| IKZF1 deletion | IKAROS transcription factor signaling | This results in activation of EBF1, MSH2, and MCL1, leading to B-cell leukemogenesis.[49] |
Genetic Diagnostic Testing Methods
- Flow cytometry for CRLF2 has been shown in some studies to be 100% concordant with FISH results[41].
- Next-generation sequencing is helpful for detecting copy number changes, single nucleotide variants, and gene fusions involving CRLF2, ABL1, ABL2, JAK2, etc.
- Gene expression profile algorithms, incorporating prediction analysis or hierarchical clustering of microarrays, provide a definitive diagnosis of B-lymphoblastic leukaemia/lymphoma with BCR::ABL1-like features.
Familial Forms
Families with certain inherited variants of GATA3 (often seen in Native-American populations) are at increased risk of B-lymphoblastic leukaemia/lymphoma with BCR::ABL1-like features.[50]
Additional Information
Clinical trial of TKI in B-lymphoblastic leukaemia/lymphoma with BCR::ABL1-like features (Clinicaltrials.gov Identifier: NCT02883049)
Clinical trial of JAK inhibitor in B-lymphoblastic leukaemia/lymphoma with BCR::ABL1-like features (Clinicaltrials.gov Identifier: NCT02723994)
Links
B-lymphoblastic leukaemia/lymphoma with BCR::ABL1-like features in Pathology Outlines
References
(use the "Cite" icon at the top of the page) (Instructions: Add each reference into the text above by clicking where you want to insert the reference, selecting the “Cite” icon at the top of the wiki page, and using the “Automatic” tab option to search by PMID to select the reference to insert. If a PMID is not available, such as for a book, please use the “Cite” icon, select “Manual” and then “Basic Form”, and include the entire reference. To insert the same reference again later in the page, select the “Cite” icon and “Re-use” to find the reference; DO NOT insert the same reference twice using the “Automatic” tab as it will be treated as two separate references. The reference list in this section will be automatically generated and sorted.)
- ↑ Mullighan, Charles G.; et al. (2009). "Deletion of IKZF1 and prognosis in acute lymphoblastic leukemia". The New England Journal of Medicine. 360 (5): 470–480. doi:10.1056/NEJMoa0808253. ISSN 1533-4406. PMC 2674612. PMID 19129520.
- ↑ Den Boer, Monique L.; et al. (2009). "A subtype of childhood acute lymphoblastic leukaemia with poor treatment outcome: a genome-wide classification study". The Lancet. Oncology. 10 (2): 125–134. doi:10.1016/S1470-2045(08)70339-5. ISSN 1474-5488. PMC 2707020. PMID 19138562.
- ↑ WHO Classification of Tumours Editorial Board. Hematolymphoid tumors. Lyon (France): International Agency for Research on Cancer; 2022. [cited 2025 NOV 05]. (WHO classification of tumors series, 5th ed.). Available from: https://tumourclassification.iarc.who.int.
- ↑ Campo, Elias; et al. (2022-09-15). "The International Consensus Classification of Mature Lymphoid Neoplasms: a report from the Clinical Advisory Committee". Blood. 140 (11): 1229–1253. doi:10.1182/blood.2022015851. ISSN 1528-0020. PMC 9479027 Check
|pmc=value (help). PMID 35653592 Check|pmid=value (help). - ↑ 5.00 5.01 5.02 5.03 5.04 5.05 5.06 5.07 5.08 5.09 5.10 5.11 Roberts, Kathryn G.; et al. (2014-09-11). "Targetable kinase-activating lesions in Ph-like acute lymphoblastic leukemia". The New England Journal of Medicine. 371 (11): 1005–1015. doi:10.1056/NEJMoa1403088. ISSN 1533-4406. PMC 4191900. PMID 25207766.
- ↑ Akkari, Yassmine M. N.; et al. (2020-05). "Evidence-based review of genomic aberrations in B-lymphoblastic leukemia/lymphoma: Report from the cancer genomics consortium working group for lymphoblastic leukemia". Cancer Genetics. 243: 52–72. doi:10.1016/j.cancergen.2020.03.001. ISSN 2210-7762. PMID 32302940 Check
|pmid=value (help). Check date values in:|date=(help) - ↑ 7.0 7.1 7.2 7.3 7.4 7.5 7.6 7.7 7.8 Reshmi, Shalini C.; et al. (2017-06-22). "Targetable kinase gene fusions in high-risk B-ALL: a study from the Children's Oncology Group". Blood. 129 (25): 3352–3361. doi:10.1182/blood-2016-12-758979. ISSN 1528-0020. PMC 5482101. PMID 28408464.
- ↑ Zaliova, Marketa; et al. (2016-09). "Characterization of leukemias with ETV6-ABL1 fusion". Haematologica. 101 (9): 1082–1093. doi:10.3324/haematol.2016.144345. ISSN 1592-8721. PMC 5060025. PMID 27229714. Check date values in:
|date=(help) - ↑ Ernst, Thomas; et al. (2011-04). "Identification of FOXP1 and SNX2 as novel ABL1 fusion partners in acute lymphoblastic leukaemia". British Journal of Haematology. 153 (1): 43–46. doi:10.1111/j.1365-2141.2010.08457.x. ISSN 1365-2141. PMID 21391972. Check date values in:
|date=(help) - ↑ Duployez, Nicolas; et al. (2016-04). "NUP214-ABL1 fusion defines a rare subtype of B-cell precursor acute lymphoblastic leukemia that could benefit from tyrosine kinase inhibitors". Haematologica. 101 (4): e133–134. doi:10.3324/haematol.2015.136499. ISSN 1592-8721. PMC 5004396. PMID 26681761. Check date values in:
|date=(help) - ↑ Collette, Y.; et al. (2015-03-13). "Drug response profiling can predict response to ponatinib in a patient with t(1;9)(q24;q34)-associated B-cell acute lymphoblastic leukemia". Blood Cancer Journal. 5 (3): e292. doi:10.1038/bcj.2015.13. ISSN 2044-5385. PMC 4382656. PMID 25768406.
- ↑ Sheng, Guangying; et al. (2017). "t(1;9)(p34;q34)/SFPQ-ABL1 Fusion in a Patient with Ph-Like Common B-Cell Acute Lymphoblastic Leukemia". Acta Haematologica. 137 (1): 40–43. doi:10.1159/000452265. ISSN 1421-9662. PMID 27894117.
- ↑ 13.00 13.01 13.02 13.03 13.04 13.05 13.06 13.07 13.08 13.09 13.10 13.11 13.12 13.13 Tasian, Sarah K.; et al. (2017-11-09). "Philadelphia chromosome-like acute lymphoblastic leukemia". Blood. 130 (19): 2064–2072. doi:10.1182/blood-2017-06-743252. ISSN 1528-0020. PMC 5680607. PMID 28972016.
- ↑ Tomita, Osamu; et al. (2014-03). "Sensitivity of SNX2-ABL1 toward tyrosine kinase inhibitors distinct from that of BCR-ABL1". Leukemia Research. 38 (3): 361–370. doi:10.1016/j.leukres.2013.11.017. ISSN 1873-5835. PMID 24367893. Check date values in:
|date=(help) - ↑ Soler, G.; et al. (2008-06). "Fusion of ZMIZ1 to ABL1 in a B-cell acute lymphoblastic leukaemia with a t(9;10)(q34;q22.3) translocation". Leukemia. 22 (6): 1278–1280. doi:10.1038/sj.leu.2405033. ISSN 1476-5551. PMID 18007576. Check date values in:
|date=(help) - ↑ Raca, Gordana; et al. (2015-04). "RCSD1-ABL2 fusion resulting from a complex chromosomal rearrangement in high-risk B-cell acute lymphoblastic leukemia". Leukemia & Lymphoma. 56 (4): 1145–1147. doi:10.3109/10428194.2014.951851. ISSN 1029-2403. PMID 25098428. Check date values in:
|date=(help) - ↑ Tran, Thai Hoa; et al. (2018-03-13). "Prognostic impact of kinase-activating fusions and IKZF1 deletions in pediatric high-risk B-lineage acute lymphoblastic leukemia". Blood Advances. 2 (5): 529–533. doi:10.1182/bloodadvances.2017014704. ISSN 2473-9537. PMC 5851421. PMID 29507076.
- ↑ 18.0 18.1 Jain, Nitin; et al. (2017-02-02). "Ph-like acute lymphoblastic leukemia: a high-risk subtype in adults". Blood. 129 (5): 572–581. doi:10.1182/blood-2016-07-726588. ISSN 1528-0020. PMC 5290985. PMID 27919910.
- ↑ Gu, Zhaohui; et al. (2016-11-08). "Genomic analyses identify recurrent MEF2D fusions in acute lymphoblastic leukaemia". Nature Communications. 7: 13331. doi:10.1038/ncomms13331. ISSN 2041-1723. PMC 5105166. PMID 27824051.
- ↑ 20.0 20.1 20.2 Iacobucci, Ilaria; et al. (2016-02-08). "Truncating Erythropoietin Receptor Rearrangements in Acute Lymphoblastic Leukemia". Cancer Cell. 29 (2): 186–200. doi:10.1016/j.ccell.2015.12.013. ISSN 1878-3686. PMC 4750652. PMID 26859458.
- ↑ Zhang, Qi; et al. (2018-01-30). "Inhibition of mTORC1/C2 signaling improves anti-leukemia efficacy of JAK/STAT blockade in CRLF2 rearranged and/or JAK driven Philadelphia chromosome-like acute B-cell lymphoblastic leukemia". Oncotarget. 9 (8): 8027–8041. doi:10.18632/oncotarget.24261. ISSN 1949-2553. PMC 5814279. PMID 29487712.
- ↑ Griesinger, Frank; et al. (2005-11). "A BCR-JAK2 fusion gene as the result of a t(9;22)(p24;q11.2) translocation in a patient with a clinically typical chronic myeloid leukemia". Genes, Chromosomes & Cancer. 44 (3): 329–333. doi:10.1002/gcc.20235. ISSN 1045-2257. PMID 16001431. Check date values in:
|date=(help) - ↑ 23.0 23.1 Roberts, Kathryn G.; et al. (2017-09-12). "Oncogenic role and therapeutic targeting of ABL-class and JAK-STAT activating kinase alterations in Ph-like ALL". Blood Advances. 1 (20): 1657–1671. doi:10.1182/bloodadvances.2017011296. ISSN 2473-9529. PMC 5728345. PMID 29296813.
- ↑ Zhou, Min-hang; et al. (2012-08). "Detection of ETV6 gene rearrangements in adult acute lymphoblastic leukemia". Annals of Hematology. 91 (8): 1235–1243. doi:10.1007/s00277-012-1431-4. ISSN 1432-0584. PMID 22373549. Check date values in:
|date=(help) - ↑ Schwaller, Jurg (2012-12). "Modeling ETV6-JAK2-induced leukemia: insights from the zebrafish". Haematologica. 97 (12): 1783–1785. doi:10.3324/haematol.2012.080754. ISSN 1592-8721. PMC 3590083. PMID 23204479. Check date values in:
|date=(help) - ↑ Ding, Yang Y.; et al. (2018-09). "Clinical efficacy of ruxolitinib and chemotherapy in a child with Philadelphia chromosome-like acute lymphoblastic leukemia with GOLGA5-JAK2 fusion and induction failure". Haematologica. 103 (9): e427–e431. doi:10.3324/haematol.2018.192088. ISSN 1592-8721. PMC 6119161. PMID 29773603. Check date values in:
|date=(help) - ↑ 27.0 27.1 27.2 27.3 Roberts, Kathryn G.; et al. (2017-02). "High Frequency and Poor Outcome of Philadelphia Chromosome-Like Acute Lymphoblastic Leukemia in Adults". Journal of Clinical Oncology: Official Journal of the American Society of Clinical Oncology. 35 (4): 394–401. doi:10.1200/JCO.2016.69.0073. ISSN 1527-7755. PMC 5455698. PMID 27870571. Check date values in:
|date=(help) - ↑ Yano, Mio; et al. (2015-12). "Identification of novel kinase fusion transcripts in paediatric B cell precursor acute lymphoblastic leukaemia with IKZF1 deletion". British Journal of Haematology. 171 (5): 813–817. doi:10.1111/bjh.13757. ISSN 1365-2141. PMID 26404892. Check date values in:
|date=(help) - ↑ Schinnerl, Dagmar; et al. (2015-02-19). "The role of the Janus-faced transcription factor PAX5-JAK2 in acute lymphoblastic leukemia". Blood. 125 (8): 1282–1291. doi:10.1182/blood-2014-04-570960. ISSN 1528-0020. PMC 4375719. PMID 25515960.
- ↑ Kawamura, Machiko; et al. (2015-07). "Identification of SPAG9 as a novel JAK2 fusion partner gene in pediatric acute lymphoblastic leukemia with t(9;17)(p24;q21)". Genes, Chromosomes & Cancer. 54 (7): 401–408. doi:10.1002/gcc.22251. ISSN 1098-2264. PMID 25951811. Check date values in:
|date=(help) - ↑ Poitras, Jennifer L.; et al. (2008-10). "Novel SSBP2-JAK2 fusion gene resulting from a t(5;9)(q14.1;p24.1) in pre-B acute lymphocytic leukemia". Genes, Chromosomes & Cancer. 47 (10): 884–889. doi:10.1002/gcc.20585. ISSN 1098-2264. PMID 18618714. Check date values in:
|date=(help) - ↑ Roberts, Kathryn G.; et al. (2012-08-14). "Genetic alterations activating kinase and cytokine receptor signaling in high-risk acute lymphoblastic leukemia". Cancer Cell. 22 (2): 153–166. doi:10.1016/j.ccr.2012.06.005. ISSN 1878-3686. PMC 3422513. PMID 22897847.
- ↑ Steeghs, Elisabeth M. P.; et al. (2017-10-27). "JAK2 aberrations in childhood B-cell precursor acute lymphoblastic leukemia". Oncotarget. 8 (52): 89923–89938. doi:10.18632/oncotarget.21027. ISSN 1949-2553. PMC 5685720. PMID 29163799.
- ↑ Kobayashi, Kenichiro; et al. (2014-06). "ATF7IP as a novel PDGFRB fusion partner in acute lymphoblastic leukaemia in children". British Journal of Haematology. 165 (6): 836–841. doi:10.1111/bjh.12834. ISSN 1365-2141. PMID 24628626. Check date values in:
|date=(help) - ↑ Ishibashi, Takeshi; et al. (2016-03). "Ph-like ALL-related novel fusion kinase ATF7IP-PDGFRB exhibits high sensitivity to tyrosine kinase inhibitors in murine cells". Experimental Hematology. 44 (3): 177–188.e5. doi:10.1016/j.exphem.2015.11.009. ISSN 1873-2399. PMID 26703895. Check date values in:
|date=(help) - ↑ Zhang, Ge; et al. (2017-11-14). "Acute Lymphoblastic Leukemia Patient with Variant ATF7IP/PDGFRB Fusion and Favorable Response to Tyrosine Kinase Inhibitor Treatment: A Case Report". The American Journal of Case Reports. 18: 1204–1208. doi:10.12659/ajcr.906300. ISSN 1941-5923. PMC 5700447. PMID 29133777.
- ↑ Schwab, Claire; et al. (2016-05-05). "EBF1-PDGFRB fusion in pediatric B-cell precursor acute lymphoblastic leukemia (BCP-ALL): genetic profile and clinical implications". Blood. 127 (18): 2214–2218. doi:10.1182/blood-2015-09-670166. ISSN 1528-0020. PMID 26872634.
- ↑ 38.0 38.1 38.2 38.3 38.4 38.5 Boer, Judith M.; et al. (2015-09). "BCR-ABL1-like cases in pediatric acute lymphoblastic leukemia: a comparison between DCOG/Erasmus MC and COG/St. Jude signatures". Haematologica. 100 (9): e354–357. doi:10.3324/haematol.2015.124941. ISSN 1592-8721. PMC 4800707. PMID 26045294. Check date values in:
|date=(help) - ↑ Boer, Judith M.; et al. (2015). "BCR-ABL1-like cases in pediatric acute lymphoblastic leukemia: a comparison between DCOG/Erasmus MC and COG/St. Jude signatures". Haematologica. 100 (9): e354–357. doi:10.3324/haematol.2015.124941. ISSN 1592-8721. PMC 4800707. PMID 26045294.
- ↑ van der Veer, Arian; et al. (2013-10-10). "Independent prognostic value of BCR-ABL1-like signature and IKZF1 deletion, but not high CRLF2 expression, in children with B-cell precursor ALL". Blood. 122 (15): 2622–2629. doi:10.1182/blood-2012-10-462358. ISSN 1528-0020. PMC 3795461. PMID 23974192.
- ↑ 41.0 41.1 Konoplev, Sergej; et al. (2017). "CRLF2-Positive B-Cell Acute Lymphoblastic Leukemia in Adult Patients: A Single-Institution Experience". American Journal of Clinical Pathology. 147 (4): 357–363. doi:10.1093/ajcp/aqx005. ISSN 1943-7722. PMID 28340183.
- ↑ Koleilat, Alaa; et al. (2022-12). "Characterization of unusual iAMP21 B-lymphoblastic leukemia (iAMP21-ALL) from the Mayo Clinic and Children's Oncology Group". Genes, Chromosomes & Cancer. 61 (12): 710–719. doi:10.1002/gcc.23084. ISSN 1098-2264. PMC 9549522 Check
|pmc=value (help). PMID 35771717 Check|pmid=value (help). Check date values in:|date=(help) - ↑ Yoda, Akinori; et al. (2010-01-05). "Functional screening identifies CRLF2 in precursor B-cell acute lymphoblastic leukemia". Proceedings of the National Academy of Sciences of the United States of America. 107 (1): 252–257. doi:10.1073/pnas.0911726107. ISSN 1091-6490. PMC 2806782. PMID 20018760.
- ↑ Goulart, Hannah; et al. (2025-11). "A Phase I/II Trial of Ruxolitinib with Chemotherapy for Patients with Relapsed and/or Refractory Philadelphia-like Acute Lymphoblastic Leukemia". Clinical Lymphoma, Myeloma & Leukemia. 25 (11): 800–807. doi:10.1016/j.clml.2025.05.013. ISSN 2152-2669. PMID 40500616 Check
|pmid=value (help). Check date values in:|date=(help) - ↑ 45.0 45.1 Quesada A, Reynolds M, Jorgensen JL, et al. Cytokine receptor-like factor 2 (CRLF2) expression in precursor B-lymphoblastic leukemia. International Clinical Cytometry Society e-Newsletter. 2014;5(1).
- ↑ Jain, Sarika; et al. (2020-02). "BCR-ABL1-like B-Acute Lymphoblastic Leukemia/Lymphoma: A Comprehensive Review". Archives of Pathology & Laboratory Medicine. 144 (2): 150–155. doi:10.5858/arpa.2019-0194-RA. ISSN 1543-2165. PMID 31644323. Check date values in:
|date=(help) - ↑ Lee, Jae Wook; et al. (2020-07). "High incidence of RAS pathway mutations among sentinel genetic lesions of Korean pediatric BCR-ABL1-like acute lymphoblastic leukemia". Cancer Medicine. 9 (13): 4632–4639. doi:10.1002/cam4.3099. ISSN 2045-7634. PMC 7333828 Check
|pmc=value (help). PMID 32378810 Check|pmid=value (help). Check date values in:|date=(help) - ↑ Senapati, Jayastu; et al. (2023-05). "Philadelphia-Like Genetic Rearrangements in Adults With B-Cell ALL: Refractoriness to Chemotherapy and Response to Tyrosine Kinase Inhibitor in ABL Class Rearrangements". JCO precision oncology. 7: e2200707. doi:10.1200/PO.22.00707. ISSN 2473-4284. PMC 10309573 Check
|pmc=value (help). PMID 37196217 Check|pmid=value (help). Check date values in:|date=(help) - ↑ van der Veer, Arian; et al. (2013). "Independent prognostic value of BCR-ABL1-like signature and IKZF1 deletion, but not high CRLF2 expression, in children with B-cell precursor ALL". Blood. 122 (15): 2622–2629. doi:10.1182/blood-2012-10-462358. ISSN 1528-0020. PMC 3795461. PMID 23974192.
- ↑ Perez-Andreu, Virginia; et al. (2013). "Inherited GATA3 variants are associated with Ph-like childhood acute lymphoblastic leukemia and risk of relapse". Nature Genetics. 45 (12): 1494–1498. doi:10.1038/ng.2803. ISSN 1546-1718. PMC 4039076. PMID 24141364.
Notes
*Primary authors will typically be those that initially create and complete the content of a page. If a subsequent user modifies the content and feels the effort put forth is of high enough significance to warrant listing in the authorship section, please contact the Associate Editor or other CCGA representative. When pages have a major update, the new author will be acknowledged at the beginning of the page, and those who contributed previously will be acknowledged below as a prior author.
Prior Author(s): Fabiola Quintero-Rivera, MD
*Citation of this Page: “B-lymphoblastic leukaemia/lymphoma with BCR::ABL1-like features”. Compendium of Cancer Genome Aberrations (CCGA), Cancer Genomics Consortium (CGC), updated 11/6/2025, https://ccga.io/index.php/HAEM5:B-lymphoblastic_leukaemia/lymphoma_with_BCR::ABL1-like_features.