HAEM5:Acute myeloid leukaemia with CBFB::MYH11 fusion: Difference between revisions
| [checked revision] | [checked revision] |
Bailey.Glen (talk | contribs) No edit summary |
Bailey.Glen (talk | contribs) No edit summary |
||
| Line 1: | Line 1: | ||
{{DISPLAYTITLE:Acute myeloid leukaemia with CBFB::MYH11 fusion}} | {{DISPLAYTITLE:Acute myeloid leukaemia with CBFB::MYH11 fusion}} | ||
[[HAEM5:Table_of_Contents|Haematolymphoid Tumours (5th ed.)]] | [[HAEM5:Table_of_Contents|Haematolymphoid Tumours (WHO Classification, 5th ed.)]] | ||
{{Under Construction}} | {{Under Construction}} | ||
| Line 7: | Line 7: | ||
}}</blockquote> | }}</blockquote> | ||
<span style="color:#0070C0">(General Instructions – The main focus of these pages is the clinically significant genetic alterations in each disease type. Use [https://www.genenames.org/ <u>HUGO-approved gene names and symbols</u>] (italicized when appropriate), [https://varnomen.hgvs.org/ HGVS-based nomenclature for variants], as well as generic names of drugs and testing platforms or assays if applicable. Please complete tables whenever possible and do not delete them (add N/A if not applicable in the table and delete the examples). Please do not delete or alter the section headings. The use of bullet points alongside short blocks of text rather than only large paragraphs is encouraged. Additional instructions below in italicized blue text should not be included in the final page content. Please also see </span><u>[[Author_Instructions]]</u><span style="color:#0070C0"> and [[Frequently Asked Questions (FAQs)|<u>FAQs</u>]] as well as contact your [[Leadership|<u>Associate Editor</u>]] or [mailto:CCGA@cancergenomics.org <u>Technical Support</u>])</span> | <span style="color:#0070C0">(General Instructions – The main focus of these pages is the clinically significant genetic alterations in each disease type. Use [https://www.genenames.org/ <u>HUGO-approved gene names and symbols</u>] (italicized when appropriate), [https://varnomen.hgvs.org/ HGVS-based nomenclature for variants], as well as generic names of drugs and testing platforms or assays if applicable. Please complete tables whenever possible and do not delete them (add N/A if not applicable in the table and delete the examples); to add (or move) a row or column to a table, click nearby within the table and select the > symbol that appears to be given options. Please do not delete or alter the section headings. The use of bullet points alongside short blocks of text rather than only large paragraphs is encouraged. Additional instructions below in italicized blue text should not be included in the final page content. Please also see </span><u>[[Author_Instructions]]</u><span style="color:#0070C0"> and [[Frequently Asked Questions (FAQs)|<u>FAQs</u>]] as well as contact your [[Leadership|<u>Associate Editor</u>]] or [mailto:CCGA@cancergenomics.org <u>Technical Support</u>])</span> | ||
==Primary Author(s)*== | ==Primary Author(s)*== | ||
| Line 204: | Line 204: | ||
==Characteristic Chromosomal Patterns== | ==Characteristic Chromosomal Patterns== | ||
Put your text here <span style="color:#0070C0">(''EXAMPLE PATTERNS: hyperdiploid; gain of odd number chromosomes including typically chromosome 1, 3, 5, 7, 11, and 17; co-deletion of 1p and 19q; complex karyotypes without characteristic genetic findings; chromothripsis'')</span> | Put your text here <span style="color:#0070C0">(''EXAMPLE PATTERNS: hyperdiploid; gain of odd number chromosomes including typically chromosome 1, 3, 5, 7, 11, and 17; co-deletion of 1p and 19q; complex karyotypes without characteristic genetic findings; chromothripsis. Do not delete table.'')</span> | ||
{| class="wikitable sortable" | {| class="wikitable sortable" | ||
| Line 214: | Line 214: | ||
!Notes | !Notes | ||
|- | |- | ||
|EXAMPLE | |<span class="blue-text">EXAMPLE:</span> | ||
Co-deletion of 1p and 18q | Co-deletion of 1p and 18q | ||
| Line 220: | Line 220: | ||
|No | |No | ||
|No | |No | ||
|EXAMPLE: | |<span class="blue-text">EXAMPLE:</span> | ||
See chromosomal rearrangements table as this pattern is due to an unbalanced derivative translocation associated with oligodendroglioma (add reference). | See chromosomal rearrangements table as this pattern is due to an unbalanced derivative translocation associated with oligodendroglioma (add reference). | ||
|} | |} | ||
<blockquote class='blockedit'>{{Box-round|title= | <blockquote class='blockedit'>{{Box-round|title=HAEM5 Conversion Notes|The content below was from the previous version of the page. Please incorporate above.}} | ||
{| class="wikitable sortable" | {| class="wikitable sortable" | ||
!Chromosomal Pattern | !Chromosomal Pattern | ||
| Line 278: | Line 278: | ||
==Genes and Main Pathways Involved== | ==Genes and Main Pathways Involved== | ||
Put your text here and fill in the table <span style="color:#0070C0">(''Instructions: Can include references in the table.'')</span> | Put your text here and fill in the table <span style="color:#0070C0">(''Instructions: Can include references in the table. Do not delete table.'')</span> | ||
{| class="wikitable sortable" | {| class="wikitable sortable" | ||
|- | |- | ||
!Gene; Genetic Alteration!!Pathway!!Pathophysiologic Outcome | !Gene; Genetic Alteration!!Pathway!!Pathophysiologic Outcome | ||
|- | |- | ||
|EXAMPLE: BRAF and MAP2K1; Activating mutations | |<span class="blue-text">EXAMPLE:</span> BRAF and MAP2K1; Activating mutations | ||
|EXAMPLE: MAPK signaling | |<span class="blue-text">EXAMPLE:</span> MAPK signaling | ||
|EXAMPLE: Increased cell growth and proliferation | |<span class="blue-text">EXAMPLE:</span> Increased cell growth and proliferation | ||
|- | |- | ||
|EXAMPLE: CDKN2A; Inactivating mutations | |<span class="blue-text">EXAMPLE:</span> CDKN2A; Inactivating mutations | ||
|EXAMPLE: Cell cycle regulation | |<span class="blue-text">EXAMPLE:</span> Cell cycle regulation | ||
|EXAMPLE: Unregulated cell division | |<span class="blue-text">EXAMPLE:</span> Unregulated cell division | ||
|- | |- | ||
|EXAMPLE: KMT2C and ARID1A; Inactivating mutations | |<span class="blue-text">EXAMPLE:</span> KMT2C and ARID1A; Inactivating mutations | ||
|EXAMPLE: Histone modification, chromatin remodeling | |<span class="blue-text">EXAMPLE:</span> Histone modification, chromatin remodeling | ||
|EXAMPLE: Abnormal gene expression program | |<span class="blue-text">EXAMPLE:</span> Abnormal gene expression program | ||
|} | |} | ||
<blockquote class='blockedit'>{{Box-round|title= | <blockquote class='blockedit'>{{Box-round|title=HAEM5 Conversion Notes|The content below was from the previous version of the page. Please incorporate above.}} | ||
''CBFB'' (core binding factor β) on 16q22 is transcribed from centromere to telomere. It codes for CBFβ, a subunit of the transcription factor complex core binding factor. CBFβ by itself does not contain any DNA binding motif or transcriptional activation domain, but forms a dimer with ''CBFa'' (''RUNX1'') which is a transcription factor . ''MYH11'' (smooth muscle myosin heavy-chain gene) on 16p13.1 is transcribed from centromere to telomere, contains a N-term ATPase head responsible for actin binding and mechanical movement, and a C-terminus long repeat of coil-coil domain to facilitate filament aggregates; member of the myosin II family. | ''CBFB'' (core binding factor β) on 16q22 is transcribed from centromere to telomere. It codes for CBFβ, a subunit of the transcription factor complex core binding factor. CBFβ by itself does not contain any DNA binding motif or transcriptional activation domain, but forms a dimer with ''CBFa'' (''RUNX1'') which is a transcription factor . ''MYH11'' (smooth muscle myosin heavy-chain gene) on 16p13.1 is transcribed from centromere to telomere, contains a N-term ATPase head responsible for actin binding and mechanical movement, and a C-terminus long repeat of coil-coil domain to facilitate filament aggregates; member of the myosin II family. | ||
Revision as of 11:28, 1 September 2024
Haematolymphoid Tumours (WHO Classification, 5th ed.)
| This page is under construction |
editHAEM5 Conversion NotesThis page was converted to the new template on 2023-12-07. The original page can be found at HAEM4:Acute Myeloid Leukemia (AML) with inv(16)(p13.1q22) or t(16;16)(p13.1;q22); CBFB-MYH11.
(General Instructions – The main focus of these pages is the clinically significant genetic alterations in each disease type. Use HUGO-approved gene names and symbols (italicized when appropriate), HGVS-based nomenclature for variants, as well as generic names of drugs and testing platforms or assays if applicable. Please complete tables whenever possible and do not delete them (add N/A if not applicable in the table and delete the examples); to add (or move) a row or column to a table, click nearby within the table and select the > symbol that appears to be given options. Please do not delete or alter the section headings. The use of bullet points alongside short blocks of text rather than only large paragraphs is encouraged. Additional instructions below in italicized blue text should not be included in the final page content. Please also see Author_Instructions and FAQs as well as contact your Associate Editor or Technical Support)
Primary Author(s)*
Christine Bryke, MD
Beth Israel Deaconess Medical Center, Boston, MA
Cancer Category / Type
Acute myeloid leukemia
Cancer Sub-Classification / Subtype
Acute myeloid leukaemia (AML) with inv(16)(p13.1q22) or t(16;16)(p13.1;q22) resulting in CBFB-MYH11 fusion
Definition / Description of Disease
Acute myeloid leukemia (AML) with inv(16)(p13.1q22) or t(16;16)(p13.1;q22) resulting in CBFB-MYH11 gene fusion is a subtype of AML with granulocytic and monocytic differentiation and abnormal bone marrow eosinophils. In the 2016 revision to the World Health Organization (WHO) classification of myeloid neoplasms and acute leukemia, it is in the group of AML with recurrent genetic abnormalities[1]. In the older French-American-British (FAB) classification system, inv(16)(p13.1q22) or t(16;16)(p13.1;q22) positive AML belonged to the acute myelomonocytic leukemia with abnormal bone marrow eosinophils M4e category. inv(16)(p13.1q22) or t(16;16)(p13.1;q22) can occasionally be seen in therapy-related AML and in blast phase of chronic myelogenous leukemia. In those instances, the leukemia is not classified in the group of AML with recurrent genetic abnormalities due to clinical and prognostic differences.
- inv(16)(p13.1q22), a pericentric inversion of chromosome 16, and the less common t(16;16)(p13.1;q22), a translocation involving the short arm of one chromosome 16 and the long arm of the other chromosome 16, define a distinctive cytogenetic subtype of acute myeloid leukemia. Both of these chromosome rearrangements result in the CBFB-MYH11 gene fusion.
- Occurs in all age groups but predominantly in younger patients.
- Myeloid sarcoma may be present at initial diagnosis or at relapse and in some patients may constitute the only evidence of relapse.
- inv(16)(p13.1q22) is a subtle chromosome rearrangement that may not be appreciated when chromosome morphology is suboptimal.
- Cases with a cryptic CBFB-MYH11 gene rearrangement are possible.
- A deletion within 16p13 accompanying a rearrangement of 16p13 and 16q22 occurs in 20% of cases[2][3].
- Loss of 3’ CBFB occurs in 4-8% of cases of CBFB rearranged AML. Of these cases, 69-73% are positive for CBFB-MYH11 fusion by RT-PCR [4][5][6][7].
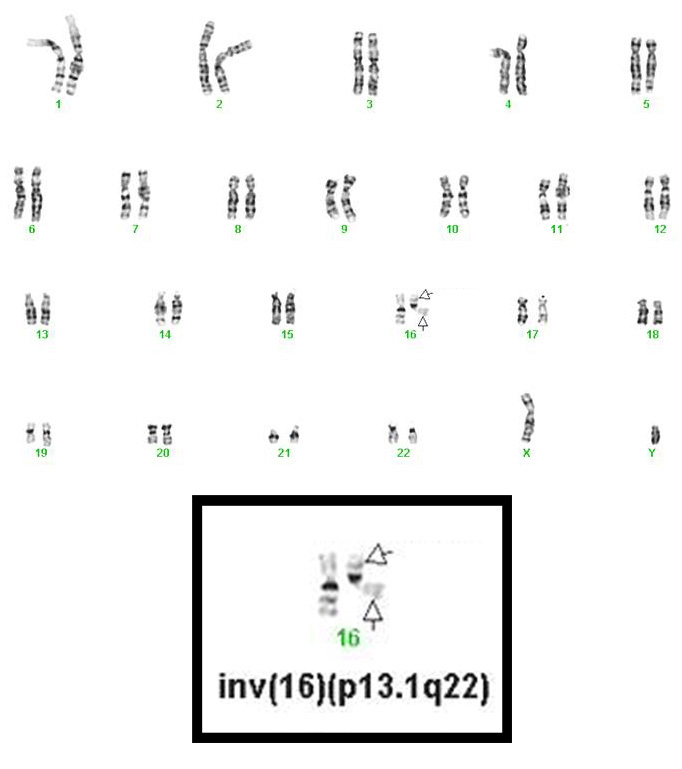
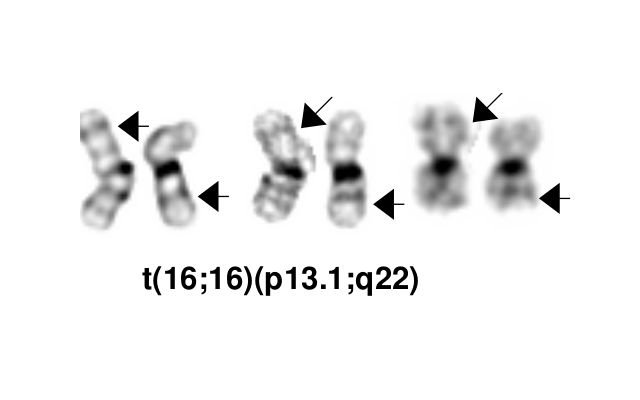
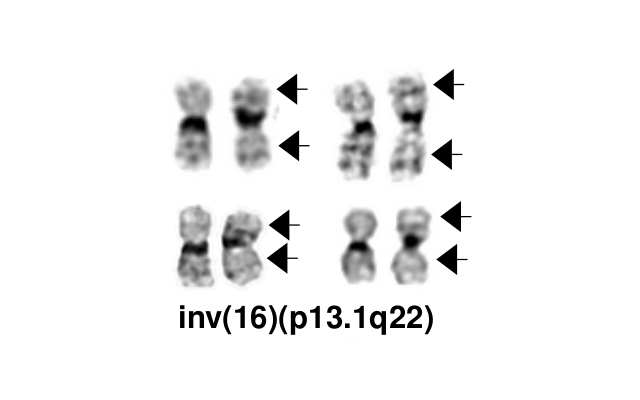

Synonyms / Terminology
Acute myeloid leukaemia (AML) with inv(16)(p13.1q22); AML, t(16;16)(p13.1;q22);AML,CBFB-MYH11; Acute myelomonocytic leukemia with abnormal eosinophils; FAB classification M4Eo
Epidemiology / Prevalence
Approximately 5-8% of AML cases have inv(16)(p13.1q22) or t(16;16)(p13.1;q22). More common in males and younger adults.
Clinical Features
| Signs and Symptoms | May present as myeloid sarcoma |
| Laboratory Findings | Abnormal eosinophil component (bone marrow)
Leukocytosis |
Sites of Involvement
Bone marrow
Morphologic Features
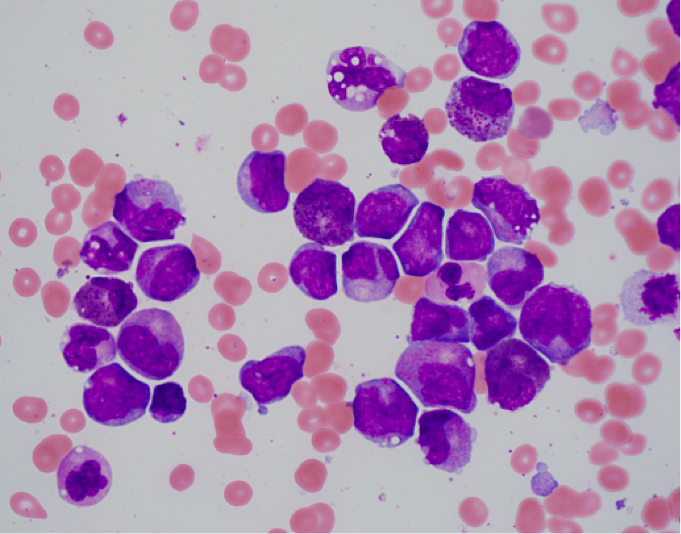
- Bone marrow shows the usual morphologic features of acute myelomonocytic leukemia.
- Eosinophils at all stages of maturation are usually increased.
- Eosinophilic granules at the promyelocyte and myelocyte stages are larger than usual and often have a purple-violet color. They may be so dense as to obscure the nucleus.
- Mature eosinophils may occasionally show nuclear hyposegmentation.
- Abnormal eosinophils are not usually seen in the peripheral blood.
- Auer rods may be present in myeloblasts.
- Occasional cases do not have eosinophilia or exhibit only myeloid maturation without a monocytic component or only monocytic maturation.
Immunophenotype
- Naphthol-ASD-chloroacetate esterase reaction, which is negative in normal eosinophils, is faintly positive in the abnormal eosinophils.
- ≥3% of blasts show myeloperoxidase activity.
- Monoblasts and promonoblasts usually show non-specific esterase activity.
Often a complex immunophenotype with multiple blast populations is seen in:
- immature blasts with high CD34 and CD117 expression
- populations with granulocytic differentiation positive for CD13, CD33, CD15, CD65 and MPO
- populations with monocytic differentiation, positive for CD14, CD4, CD11c, CD11b, CD11c, CD64, CD36 and lysozyme
| Finding | Marker |
|---|---|
| Positive (granulocytic differentiation) | CD13, CD33, CD15, CD65 and MPO |
| Positive (monocytic differentiation) | CD14, CD4, CD11c, CD11b, CD11c, CD64, CD36 and lysozyme |
| Positive (subset) | |
| Negative (universal) | |
| Negative (subset) |
Chromosomal Rearrangements (Gene Fusions)
| Chromosomal Rearrangement | Genes in Fusion (5’ or 3’ Segments) | Pathogenic Derivative | Prevalence | Diagnostic Significance (Yes, No or Unknown) | Prognostic Significance (Yes, No or Unknown) | Therapeutic Significance (Yes, No or Unknown) | Notes |
|---|---|---|---|---|---|---|---|
| inv(16)(p13.1q22) or t(16;16)(p13.1;q22) | 5'CBFB / 3'MYH11 | der(16) | 5-8% of AML | Yes | Yes | No | Diagnostic
Per 2022 International Consensus Classification of Myeloid Neoplasms and Acute Leukemias guidelines, blast count needs to be at least 10% or more [8]
|
Individual Region Genomic Gain / Loss / LOH
inv(16)(p13.1q22) or t(16;16)(p13.1;q22) is the sole chromosome aberration in approximately 60% of cases.
| Chr # | Gain / Loss / Amp / LOH | Minimal Region Genomic Coordinates [Genome Build] | Minimal Region Cytoband | Diagnostic Significance (Yes, No or Unknown) | Prognostic Significance (Yes, No or Unknown) | Therapeutic Significance (Yes, No or Unknown) | Notes |
|---|---|---|---|---|---|---|---|
| 8 | Gain | chr8:1-145,138,636 [hg38] | chr8 | No | No | No | Frequently observed. Does not adversely affect the favorable prognosis associated with inv(16)(p13.1q22) or t(16;16)(p13.1;q22) unless a KIT gene mutation is also present (see below) |
| 22 | Gain | chr22:1-50,818,468 [hg38] | chr22 | No | No | No | Frequently observed. Does not adversely affect the favorable prognosis associated with inv(16)(p13.1q22) or t(16;16)(p13.1;q22) unless a KIT gene mutation is also present (see below) |
| 21 | Gain | chr21:1-46,709,983 [hg38] | chr21 | No | No | No | Occasionally observed. Does not adversely affect the favorable prognosis associated with inv(16)(p13.1q22) or t(16;16)(p13.1;q22) unless a KIT gene mutation is also present (see below) |
| 7 | Loss | chr7:60,100,001-159,345,973 [hg38] | chr7q | No | No | No | Occasionally observed. Does not adversely affect the favorable prognosis associated with inv(16)(p13.1q22) or t(16;16)(p13.1;q22) unless a KIT gene mutation is also present (see below) |
Characteristic Chromosomal Patterns
Put your text here (EXAMPLE PATTERNS: hyperdiploid; gain of odd number chromosomes including typically chromosome 1, 3, 5, 7, 11, and 17; co-deletion of 1p and 19q; complex karyotypes without characteristic genetic findings; chromothripsis. Do not delete table.)
| Chromosomal Pattern | Diagnostic Significance (Yes, No or Unknown) | Prognostic Significance (Yes, No or Unknown) | Therapeutic Significance (Yes, No or Unknown) | Notes |
|---|---|---|---|---|
| EXAMPLE:
Co-deletion of 1p and 18q |
Yes | No | No | EXAMPLE:
See chromosomal rearrangements table as this pattern is due to an unbalanced derivative translocation associated with oligodendroglioma (add reference). |
editHAEM5 Conversion NotesThe content below was from the previous version of the page. Please incorporate above.
Chromosomal Pattern Diagnostic Significance (Yes, No or Unknown) Prognostic Significance (Yes, No or Unknown) Therapeutic Significance (Yes, No or Unknown) Notes Not Applicable N/A N/A N/A N/A
Gene Mutations (SNV / INDEL)
AML with inv(16)(p13.1q22) or t(16;16)(p13.1;q22) has genetic heterogeneity at the molecular level. Several genes have been identified that are frequently mutated in this subset of AML, some of them adversely affecting prognosis.
| Gene; Genetic Alteration | Presumed Mechanism (Tumor Suppressor Gene [TSG] / Oncogene / Other) | Prevalence (COSMIC / TCGA / Other) | Concomitant Mutations | Mutually Exclusive Mutations | Diagnostic Significance (Yes, No or Unknown) | Prognostic Significance (Yes, No or Unknown) | Therapeutic Significance (Yes, No or Unknown) | Notes |
|---|---|---|---|---|---|---|---|---|
| KIT; Gain of Function mutations in exons 8 or 17 | Proto-Oncogene | 30%[9] | N/A | N/A | No | Yes | The KIT gene on 4q12 encodes a transmembrane glycoprotein that activates signaling pathways involved in cellular proliferation, differentiation and survival. Associated with a high risk of relapse in cases of AML with inv(16)(p13.1q22) or t(16;16)(p13.1;q22). Patients with KIT mutations have a significantly shorter overall survival and disease free survival periods than those without KIT mutations. Gene expression profile analysis of KIT mutation positive core binding factor AML showed dysregulation of genes belonging to the NF-κB signaling complex indicating impairment of apoptosis[12]. |
Other Mutations
Mutations in NRAS (45%), KRAS and FLT3 (FLT3-TKD and FLT3-ITD) have also been found in inv(16)(p13.1q22) or t(16;16)(p13.1;q22) AML[13]. Recently a FLT3 N676K mutation has been identified in a small subset of the disease[14][15]. In addition, TET2 mutations have been identified[16] and WT1 mutations have been found to be relatively common (13.8%) in inv(16) core-binding factor leukemia[17]. CBL mutations are less common and can be seen in ~5% of patients with inv(16) AML[18].
- Negative for ASXL2 mutation [which is common in t(8;21)].
Note: A more extensive list of mutations can be found in cBioportal (https://www.cbioportal.org/), COSMIC (https://cancer.sanger.ac.uk/cosmic), ICGC (https://dcc.icgc.org/) and/or other databases. When applicable, gene-specific pages within the CCGA site directly link to pertinent external content.
Epigenomic Alterations
None
Genes and Main Pathways Involved
Put your text here and fill in the table (Instructions: Can include references in the table. Do not delete table.)
| Gene; Genetic Alteration | Pathway | Pathophysiologic Outcome |
|---|---|---|
| EXAMPLE: BRAF and MAP2K1; Activating mutations | EXAMPLE: MAPK signaling | EXAMPLE: Increased cell growth and proliferation |
| EXAMPLE: CDKN2A; Inactivating mutations | EXAMPLE: Cell cycle regulation | EXAMPLE: Unregulated cell division |
| EXAMPLE: KMT2C and ARID1A; Inactivating mutations | EXAMPLE: Histone modification, chromatin remodeling | EXAMPLE: Abnormal gene expression program |
editHAEM5 Conversion NotesThe content below was from the previous version of the page. Please incorporate above.CBFB (core binding factor β) on 16q22 is transcribed from centromere to telomere. It codes for CBFβ, a subunit of the transcription factor complex core binding factor. CBFβ by itself does not contain any DNA binding motif or transcriptional activation domain, but forms a dimer with CBFa (RUNX1) which is a transcription factor . MYH11 (smooth muscle myosin heavy-chain gene) on 16p13.1 is transcribed from centromere to telomere, contains a N-term ATPase head responsible for actin binding and mechanical movement, and a C-terminus long repeat of coil-coil domain to facilitate filament aggregates; member of the myosin II family.
DNA: inv(16)(p13.1q22) and t(16;16)(p13.1;q22) result in the CBFB-MYH11 hybrid gene involving fusion of 5’ CBFB in intron 5 and variable breakpoints in MYH11.
mRNA: At least 8 different CBFB-MYH11 fusion transcripts have been described. A transcript with CBFB and MYH11 positions at nucleotides 495 and 1921, respectively, is found in about 90% of patients. There is no reciprocal MYH11-CBFB transcript.
Protein: A fusion protein with the first 165 (or 133 in a few cases) amino acids of the N-terminus of CBFβ, minus only 17 or 22 amino acids, fused to the tail of the MYH11 C-terminus with its multimerization domain. The breakpoint in MYH11 is variable. The fusion protein retains the ability to dimerize with RUNX1 in the heterodimeric core binding factor. The core binding factor is a transcription factor that plays an essential role in regulation of normal hematopoiesis. It is composed of a DNA-binding CBFα (RUNX1) chain and a non-DNA-binding CBFβ chain. It is likely oncogenic due to altered transcriptional regulation of normal RUNX1 target genes. Animal studies suggest that the fusion proteins alone are not able to induce leukemia and that additional genetic alterations are required for leukemogenic transformation[19].
Genetic Diagnostic Testing Methods
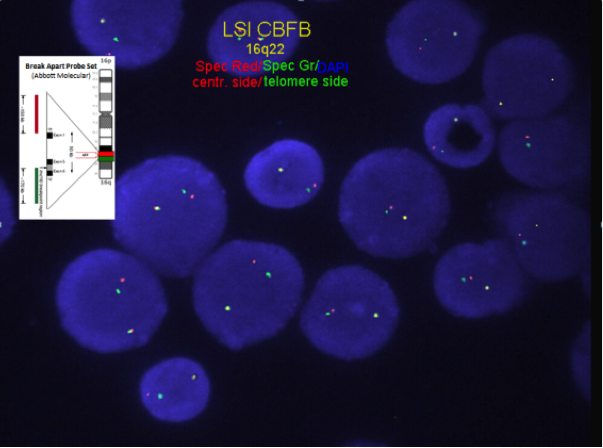
Chromosome analysis for inversion or translocation; FISH or RT-PCR for CBFB-MYH11 gene fusion rearrangement.
FISH or RT-PCR for CBFB-MYH11 is needed when chromosome morphology is suboptimal and for cytogenetically cryptic cases with typical bone marrow morphology and immunophenotype.
- RT-PCR for CBFB-MYH11 is employed for minimal residual disease detection. The attainment of optimal PCR response (OPR) of <0.1% PCR transcripts (at a level of detection of 10-4 ) after induction (C1) and <0.01% during or following consolidation has led to improved relapse-free survival (RFS)[20][21]
- In KIT mutation positive cases detection of the KIT mutation should not be used for assessment minimal residual disease because KIT mutations are frequently lost at relapse[18].
Familial Forms
None
Additional Information
None
Links
http://www.uptodate.com/contents/cytogenetics-in-acute-myeloid-leukemia
http://www.genenames.org/cgi-bin/gene_symbol_report?hgnc_id=HGNC:1539
http://www.clevelandclinicmeded.com.
References
- ↑ Arber, Daniel A.; et al. (05 19, 2016). "The 2016 revision to the World Health Organization classification of myeloid neoplasms and acute leukemia". Blood. 127 (20): 2391–2405. doi:10.1182/blood-2016-03-643544. ISSN 1528-0020. PMID 27069254. Check date values in:
|date=(help) - ↑ Marlton, P.; et al. (1995-02-01). "Molecular characterization of 16p deletions associated with inversion 16 defines the critical fusion for leukemogenesis". Blood. 85 (3): 772–779. ISSN 0006-4971. PMID 7833479.
- ↑ Martinet, D.; et al. (1997-07). "Detection of 16 p deletions by FISH in patients with inv(16) or t(16;16) and acute myeloid leukemia (AML)". Leukemia. 11 (7): 964–970. doi:10.1038/sj.leu.2400681. ISSN 0887-6924. PMID 9204976. Check date values in:
|date=(help) - ↑ Yang, Richard K.; et al. (2021-10-26). "CBFB Break-Apart FISH Testing: An Analysis of 1629 AML Cases with a Focus on Atypical Findings and Their Implications in Clinical Diagnosis and Management". Cancers. 13 (21): 5354. doi:10.3390/cancers13215354. ISSN 2072-6694. PMC 8582369 Check
|pmc=value (help). PMID 34771519 Check|pmid=value (help). - ↑ Lv, Lili; et al. (2020). "Acute myeloid leukemia with inv(16)(p13.1q22) and deletion of the 5'MYH11/3'CBFB gene fusion: a report of two cases and literature review". Molecular Cytogenetics. 13: 4. doi:10.1186/s13039-020-0474-9. ISSN 1755-8166. PMC 6990480. PMID 32015759 Check
|pmid=value (help). - ↑ Kelly, Johanna; et al. (2005-10-15). "3'CBFbeta deletion associated with inv(16) in acute myeloid leukemia". Cancer Genetics and Cytogenetics. 162 (2): 122–126. doi:10.1016/j.cancergencyto.2005.03.001. ISSN 0165-4608. PMID 16213359.
- ↑ Tang, Guilin; et al. (2022-04). "3'CBFB deletion in CBFB-rearranged acute myeloid leukemia retains morphological features associated with inv(16), but patients have higher risk of relapse and may require stem cell transplant". Annals of Hematology. 101 (4): 847–854. doi:10.1007/s00277-022-04767-1. ISSN 1432-0584. PMID 35184217 Check
|pmid=value (help). Check date values in:|date=(help) - ↑ Arber, Daniel A.; et al. (2022-09-15). "International Consensus Classification of Myeloid Neoplasms and Acute Leukemias: integrating morphologic, clinical, and genomic data". Blood. 140 (11): 1200–1228. doi:10.1182/blood.2022015850. ISSN 0006-4971. PMC PMC9479031 Check
|pmc=value (help). PMID 35767897 Check|pmid=value (help).CS1 maint: PMC format (link) - ↑ 9.0 9.1 Paschka, Peter; et al. (2006-08-20). "Adverse prognostic significance of KIT mutations in adult acute myeloid leukemia with inv(16) and t(8;21): a Cancer and Leukemia Group B Study". Journal of Clinical Oncology: Official Journal of the American Society of Clinical Oncology. 24 (24): 3904–3911. doi:10.1200/JCO.2006.06.9500. ISSN 1527-7755. PMID 16921041.
- ↑ Senapati, Jayastu; et al. (2023-03). "Common kinase mutations do not impact optimal molecular responses in core binding factor acute myeloid leukemia treated with fludarabine, cytarabine, and G‐CSF based regimens". American Journal of Hematology. 98 (3). doi:10.1002/ajh.26811. ISSN 0361-8609. Check date values in:
|date=(help) - ↑ Yoon, J.-H.; et al. (2014-12). "Identification of molecular and cytogenetic risk factors for unfavorable core-binding factor-positive adult AML with post-remission treatment outcome analysis including transplantation". Bone Marrow Transplantation. 49 (12): 1466–1474. doi:10.1038/bmt.2014.180. ISSN 1476-5365. PMID 25111512. Check date values in:
|date=(help) - ↑ 12.0 12.1 Solh, Melhem; et al. (2014-12). "Core-binding factor acute myeloid leukemia: Heterogeneity, monitoring, and therapy". American Journal of Hematology. 89 (12): 1121–1131. doi:10.1002/ajh.23821. ISSN 1096-8652. PMID 25088818. Check date values in:
|date=(help) - ↑ Paschka, Peter; et al. (2013-01-03). "Secondary genetic lesions in acute myeloid leukemia with inv(16) or t(16;16): a study of the German-Austrian AML Study Group (AMLSG)". Blood. 121 (1): 170–177. doi:10.1182/blood-2012-05-431486. ISSN 1528-0020. PMID 23115274.
- ↑ Opatz, Sabrina; et al. (2013-09-05). "Exome sequencing identifies recurring FLT3 N676K mutations in core-binding factor leukemia". Blood. 122 (10): 1761–1769. doi:10.1182/blood-2013-01-476473. ISSN 1528-0020. PMID 23878140.
- ↑ Huang, Kezhi; et al. (2016-04). "Leukemogenic potency of the novel FLT3-N676K mutant". Annals of Hematology. 95 (5): 783–791. doi:10.1007/s00277-016-2616-z. ISSN 1432-0584. PMID 26891877. Check date values in:
|date=(help) - ↑ Cher, C. Y.; et al. (07 08, 2016). "Next-generation sequencing with a myeloid gene panel in core-binding factor AML showed KIT activation loop and TET2 mutations predictive of outcome". Blood Cancer Journal. 6 (7): e442. doi:10.1038/bcj.2016.51. ISSN 2044-5385. PMC 5030377. PMID 27391574. Check date values in:
|date=(help) - ↑ Park, Sang Hyuk; et al. (2015-05). "Incidences and Prognostic Impact of c-KIT, WT1, CEBPA, and CBL Mutations, and Mutations Associated With Epigenetic Modification in Core Binding Factor Acute Myeloid Leukemia: A Multicenter Study in a Korean Population". Annals of Laboratory Medicine. 35 (3): 288–297. doi:10.3343/alm.2015.35.3.288. ISSN 2234-3814. PMC 4390696. PMID 25932436. Check date values in:
|date=(help) - ↑ 18.0 18.1 Allen, C.; et al. (2013-09). "The importance of relative mutant level for evaluating impact on outcome of KIT, FLT3 and CBL mutations in core-binding factor acute myeloid leukemia". Leukemia. 27 (9): 1891–1901. doi:10.1038/leu.2013.186. ISSN 1476-5551. PMID 23783394. Check date values in:
|date=(help) - ↑ Downing, James R. (2003-02). "The core-binding factor leukemias: lessons learned from murine models". Current Opinion in Genetics & Development. 13 (1): 48–54. doi:10.1016/s0959-437x(02)00018-7. ISSN 0959-437X. PMID 12573435. Check date values in:
|date=(help) - ↑ Boddu, Prajwal; et al. (2018-06-08). "Response kinetics and factors predicting survival in core-binding factor leukemia". Leukemia. 32 (12): 2698–2701. doi:10.1038/s41375-018-0158-1. ISSN 0887-6924.
- ↑ Yin, John A. Liu; et al. (2012-10-04). "Minimal residual disease monitoring by quantitative RT-PCR in core binding factor AML allows risk stratification and predicts relapse: results of the United Kingdom MRC AML-15 trial". Blood. 120 (14): 2826–2835. doi:10.1182/blood-2012-06-435669. ISSN 0006-4971.
Notes
*Primary authors will typically be those that initially create and complete the content of a page. If a subsequent user modifies the content and feels the effort put forth is of high enough significance to warrant listing in the authorship section, please contact the CCGA coordinators (contact information provided on the homepage). Additional global feedback or concerns are also welcome.
Edited by: Fabiola Quintero-Rivera 7/21/2018
*Citation of this Page: “Acute myeloid leukaemia with CBFB::MYH11 fusion”. Compendium of Cancer Genome Aberrations (CCGA), Cancer Genomics Consortium (CGC), updated 09/1/2024, https://ccga.io/index.php/HAEM5:Acute_myeloid_leukaemia_with_CBFB::MYH11_fusion.