B-lymphoblastic leukaemia/lymphoma with BCR::ABL1-like features
Haematolymphoid Tumours (WHO Classification, 5th ed.)
| This page is under construction |
editContent Update To WHO 5th Edition Classification Is In Process; Content Below is Based on WHO 4th Edition ClassificationThis page was converted to the new template on 2023-12-07. The original page can be found at HAEM4:B-Lymphoblastic Leukemia/Lymphoma, BCR-ABL1-Like.
(General Instructions – The focus of these pages is the clinically significant genetic alterations in each disease type. This is based on up-to-date knowledge from multiple resources such as PubMed and the WHO classification books. The CCGA is meant to be a supplemental resource to the WHO classification books; the CCGA captures in a continually updated wiki-stye manner the current genetics/genomics knowledge of each disease, which evolves more rapidly than books can be revised and published. If the same disease is described in multiple WHO classification books, the genetics-related information for that disease will be consolidated into a single main page that has this template (other pages would only contain a link to this main page). Use HUGO-approved gene names and symbols (italicized when appropriate), HGVS-based nomenclature for variants, as well as generic names of drugs and testing platforms or assays if applicable. Please complete tables whenever possible and do not delete them (add N/A if not applicable in the table and delete the examples); to add (or move) a row or column in a table, click nearby within the table and select the > symbol that appears. Please do not delete or alter the section headings. The use of bullet points alongside short blocks of text rather than only large paragraphs is encouraged. Additional instructions below in italicized blue text should not be included in the final page content. Please also see Author_Instructions and FAQs as well as contact your Associate Editor or Technical Support.)
Primary Author(s)*
Mark G. Evans, MD, Caris Life Sciences
Kilannin Krysiak, PhD, WashU Medicine
Sumire K. Kitahara, MD, Cedars-Sinai Medical Center
WHO Classification of Disease
| Structure | Disease |
|---|---|
| Book | Haematolymphoid Tumours (5th ed.) |
| Category | B-cell lymphoid proliferations and lymphomas |
| Family | Precursor B-cell neoplasms |
| Type | B-lymphoblastic leukaemias/lymphomas |
| Subtype(s) | B-lymphoblastic leukaemia/lymphoma with BCR::ABL1-like features |
Related Terminology
| Acceptable | Philadelphia-like (Ph-like) B-ALL; BCR::ABL1-like B-ALL/LBL |
| Not Recommended | N/A |
Gene Rearrangements
Table derived from Akkari et al., 2020 [1] with permission from Cancer Genetics.
| 3’ Partner | 5’ Partner | Chromosome rearrangement | Gene fusion | Visible by G-banding | References | Comment |
| ABL1
(9q34) |
CENPC1 | t(4;9)(q13;q34) | CENPC1-ABL1 | YES | [2] | Requires complex rearrangement due to incompatible orientation of genes with respect to chromosome arms |
| ETV6 | t(9;12)(q34;p13) | ETV6-ABL1 | NO | [3] | Requires complex rearrangement due to incompatible orientation of genes with respect to chromosome arms | |
| FOXP1 | t(3;9)(p13;q34) | FOXP1-ABL1 on der(3) | YES | [4] | ||
| LSM14A | t(9;19)(q34;q13.1) | LSM14A-ABL1 on der(19) | YES | [2] | ||
| NUP153 | t(6;9)(p22.3;q34) | NUP153-ABL1 on der(6) | YES | [2] | ||
| NUP214 | dup(9)(q34.1q34.1) | NUP214-ABL1 | NO | [5] | Tandem duplication (~370 kb) detectable by CMA | |
| RANBP2 | t(2;9)(q12.3;q34) | RANBP-ABL1 on der(2) | YES | [6] | ||
| RCSD1 | t(1;9)(q24.2;q34) | RCSD1-ABL1 on der(1) | YES | |||
| SFPQ | t(1;9)(p34.3;q34) | SFPQ-ABL1 on der(1) | YES | |||
| SNX1 | t(9;15)(q34;q22.3) | SNX1-ABL1 on der(15) | YES | |||
| SNX2 | t(5;9)(q23.2;q34) | SNX2-ABL1 on der(5) | YES | |||
| ZMIZ1 | t(9;10)(q34;q22.3) | ZMIZ1-ABL1 on der(10) | YES | |||
| ABL2
(1q25.2) |
PAG1 | t(1;8)(q25.2;q21.1) | PAG1-ABL2 on der(1) | YES | ||
| RCSD1 | 1q24.2q25.2 rearrangement | RCSD1-ABL2 | NO | On the same chromosome arm; however, a simple deletion cannot cause the fusion due to the orientation of genes | ||
| ZC3HAV1 | t(1;7)(q25.2;q34) | ZC3HAV1-ABL2 on der(1) | YES | |||
| CRLF2
(Xp22.3 & Yp11.3) |
IGH | t(X;14)(p22.3;q32) or
t(Y;14)(p11.3;q32) |
IGH/CRLF2 | NO | ||
| P2RY8 | del(X)(p22.3p22.3) or del(Y)(p11.3p11.3) | P2RY8-CRLF2 | NO | |||
| CSF1R
(5q32) |
MEF2D | t(1;5)(q22;q32) | MEF2D-CSF1R on der(5) | YES | ||
| SSBP2 | 5q14.1q32 rearrangement | SSBP2-CSF1R | YES | On the same chromosome arm; however, a simple deletion cannot cause the fusion due to the orientation of genes | ||
| TBL1XR1 | t(3;5)(q26.3;q32) | TBL1XR1-CSF1R on der(5) | YES | |||
| DGKH (13q14.1) | ZFAND3 | t(6;13)(p21.2;q14.1) | ZFAND3-DGKH | YES | Requires complex rearrangement due to incompatible orientation of genes with respect to chromosome arms | |
| EPOR (19p13.2) | IGH | ins(14;19)(q32;p13.2p13.2) | IGH/EPOR | Cryptic insertion | ||
| IGK | ins(2;19)(p11.2;p13.2p13.2) | IGK/EPOR | Cryptic insertion | |||
| LAIR1 | inv(19)(p13.2q13.42) | LAIR1-EPOR | NO | Inversion of chromosome 19 juxtaposes EPOR to the upstream region of LAIR1 | ||
| THADA | t(2;19)(p21;p13.2) | THADA-EPOR | YES | |||
| IL2RB (22q12.3) | MYH9 | 22q12.3 rearrangement | MYH9-IL2RB | NO | On the same chromosome arm; however, a simple deletion cannot cause the fusion due to the orientation of genes | |
| JAK2
(9p24.1) |
ATF7IP | t(9;12)(p24.1;p13.1) | ATF7IP-JAK2 on der(9) | NO | ||
| BCR | t(9;22)(p24.1;q11.2) | BCR-JAK2 | ? YES | Seen also in myeloproliferative neoplasms. Requires complex rearrangement due to incompatible orientation of genes with respect to chromosome arms | ||
| EBF1 | t(5;9)(q33.3;p24.1) | EBF1-JAK2 on der(9) | NO (SUBTLE) | |||
| ETV6 | t(9;12)(p24.1;p13.2) | ETV6-JAK2 on der(9) | NO (SUBTLE) | |||
| GOLGA5 | t(9;14)(p24.1;q32.1) | GOLGA5-JAK2 | NO (SUBTLE) | Requires complex rearrangement due to incompatible orientation of genes with respect to chromosome arms | ||
| HMBOX1 | t(8;9)(p21.1;p24.1) | HMBOX1-JAK2 on der(9) | YES | |||
| OFD1 | t(X;9)(p22.2;p24.1) | OFD1-JAK2 on der(9) | NO (SUBTLE) | |||
| PAX5 | inv(9)(p13.2p24.1) | PAX5-JAK2 | YES | An inversion is required as genes are oriented in opposite directions | ||
| PCM1 | t(8;9)(p22;p24.1) | PCM1-JAK2 on der(9) | YES (SUBTLE) | Seen also in myeloid/lymphoid neoplasms with eosinophilia | ||
| PPFIBP1 | t(9;12)(p24.1;p11.2) | PPFIBP1-JAK2 on der(9) | YES | |||
| RFX3 | inv(9)(p24.1p24.2) | RFX3-JAK2 | NO | An inversion is required as genes are oriented in opposite directions | ||
| SMU1 | inv(9)(p21.1p24.1) | SMU1-JAK2 | NO | An inversion is required as genes are oriented in opposite directions | ||
| SNX29 | t(9;16)(p24.1;p13.1) | SNX29-JAK2 on der(9) | YES | |||
| SPAG9 | t(9;17)(p24.1;q21.3) | SPAG9-JAK2 on der(9) | YES | |||
| SSBP2 | t(5;9)(q14.1;p24.1) | SSBP2-JAK2 on der(9) | YES | |||
| STRN3 | t(9;14)(p24.1;q12) | STRN3-JAK2 on der(9) | YES | |||
| TERF2 | t(9;16)(p24.1;q22.1) | TERF2-JAK2 on der(9) | YES | |||
| TPR | t(1;9)(q31.1;p24.1) | TPR-JAK2 on der(9) | YES | |||
| USP25 | t(9;21)(p24.1;q21.1) | USP25-JAK2 | ? YES | Requires complex rearrangement due to incompatible orientation of genes with respect to chromosome arms | ||
| ZBTB46 | t(9;20)(p24.1;q13.3) | ZBTB46-JAK2 on der(9) | NO | |||
| ZNF274 | t(9;19)(p24.1;q13.4) | ZNF274-JAK2 | NO | Requires complex rearrangement due to incompatible orientation of genes with respect to chromosome arms | ||
| ZNF340 | t(9;20)(p24.1;q13.3) | ZNF340-JAK2 on der(9) | NO | |||
| PDGFRA
(4q12) |
FIP1L1 | del(4)(q12q12) | FIP1L1-PDGFRA | NO | Interstitial deletion. Seen also in myeloid/lymphoid neoplasms with eosinophilia | |
| PDGFRB (5q32) | ATF7IP | t(5;12)(q32;p13.1) | ATF7IP-PDGFRB on der(5) | YES | ||
| EBF1 | del(5)(q32q33.3) | EBF1-PDGFRB | NO | Interstitial deletion | ||
| ETV6 | t(5;12)(q32;p13.2) | ETV6-PDGFRB on der(5) | YES | |||
| SNX29 | t(5;16)(q32;p13.1) | SNX29-PDGFRB on der(5) | YES | |||
| SSBP2 | t(5;5)(q14.1;q32) | SSBP2-PDGFRB | ? YES | On the same chromosome arm; however, a simple deletion cannot cause the fusion due to the orientation of genes | ||
| TNIP1 | del(5)(q32q33.1) | TNIP1-PDGFRB | NO | Interstitial deletion. Seen also in myeloid/lymphoid neoplasms with eosinophilia | ||
| ZEB2 | t(2;5)(q22.3;q32) | ZEB2-PDGFRB on der(5) | YES | |||
| ZMYND8 | t(5;20)(q32;q13.1) | ZMYND8-PDGFRB on der(5) | YES | |||
| PTK2B (8p21.2) | KDM6A | t(X;8)(p11.3;p21.2) | KDM6A-PTK2B on der(8) | YES | ||
| STAG2 | t(X;8)(q25;p21.2) | STAG2-PTK2B | YES | Requires complex rearrangement due to incompatible orientation of genes with respect to chromosome arms | ||
| TMEM2 | t(8;9)(p21.2;q21.1) | TMEM2-PTK2B on der(8) | YES | |||
| TYK2 (19p13.2) | MYB | t(6;19)(q23.3;p13.2) | MYB-TYK2 on der(6) | YES | ||
| SMARCA4 | inv(19)(p13.2p13.2) | SMARCA4-TYK2 | NO | |||
| ZNF340 | t(19;20)(p13.2;q13.3) | ZNF340-TYK2 | NO | Requires complex rearrangement due to incompatible orientation of genes with respect to chromosome arms |
Put your text here and fill in the table (Instructions: Details on clinical significance such as prognosis and other important information can be provided in the notes section. Please include references throughout the table. Do not delete the table.)
| Driver Gene | Fusion(s) and Common Partner Genes | Molecular Pathogenesis | Typical Chromosomal Alteration(s) | Prevalence -Common >20%, Recurrent 5-20% or Rare <5% (Disease) | Diagnostic, Prognostic, and Therapeutic Significance - D, P, T | Established Clinical Significance Per Guidelines - Yes or No (Source) | Clinical Relevance Details/Other Notes |
|---|---|---|---|---|---|---|---|
| EXAMPLE: ABL1 | EXAMPLE: BCR::ABL1 | EXAMPLE: The pathogenic derivative is the der(22) resulting in fusion of 5’ BCR and 3’ABL1. | EXAMPLE: t(9;22)(q34;q11.2) | EXAMPLE: Common (CML) | EXAMPLE: D, P, T | EXAMPLE: Yes (WHO, NCCN) | EXAMPLE:
The t(9;22) is diagnostic of CML in the appropriate morphology and clinical context (add reference). This fusion is responsive to targeted therapy such as Imatinib (Gleevec) (add reference). BCR::ABL1 is generally favorable in CML (add reference). |
| EXAMPLE: CIC | EXAMPLE: CIC::DUX4 | EXAMPLE: Typically, the last exon of CIC is fused to DUX4. The fusion breakpoint in CIC is usually intra-exonic and removes an inhibitory sequence, upregulating PEA3 genes downstream of CIC including ETV1, ETV4, and ETV5. | EXAMPLE: t(4;19)(q25;q13) | EXAMPLE: Common (CIC-rearranged sarcoma) | EXAMPLE: D | EXAMPLE:
DUX4 has many homologous genes; an alternate translocation in a minority of cases is t(10;19), but this is usually indistinguishable from t(4;19) by short-read sequencing (add references). | |
| EXAMPLE: ALK | EXAMPLE: ELM4::ALK
|
EXAMPLE: Fusions result in constitutive activation of the ALK tyrosine kinase. The most common ALK fusion is EML4::ALK, with breakpoints in intron 19 of ALK. At the transcript level, a variable (5’) partner gene is fused to 3’ ALK at exon 20. Rarely, ALK fusions contain exon 19 due to breakpoints in intron 18. | EXAMPLE: N/A | EXAMPLE: Rare (Lung adenocarcinoma) | EXAMPLE: T | EXAMPLE:
Both balanced and unbalanced forms are observed by FISH (add references). | |
| EXAMPLE: ABL1 | EXAMPLE: N/A | EXAMPLE: Intragenic deletion of exons 2–7 in EGFR removes the ligand-binding domain, resulting in a constitutively active tyrosine kinase with downstream activation of multiple oncogenic pathways. | EXAMPLE: N/A | EXAMPLE: Recurrent (IDH-wildtype Glioblastoma) | EXAMPLE: D, P, T | ||
editv4:Chromosomal Rearrangements (Gene Fusions)The content below was from the old template. Please incorporate above.
Tyrosine kinase-type translocations are common and involve ABL1 and other kinases (such as ABL2, EPOR, JAK2, PDGFRB, and CSF1R); more than 30 gene partners have been described. Frequently reported examples include IGH–EPOR of the t(14;19)(q32;p13)/ins(14;19)(q32;p13), EBF1–PDGFRB of the del(5)(q32q33.3), NUP214–ABL1 of the t(9;9)(q34;q34)/del(9)(q34q34), and ETV6–ABL1 of the t(9;12)(q34;p13). Other notable fusions are BCR–JAK2, PAX5–JAK2, STRN3–JAK2, RANBP2–ABL1, RCSD1–ABL1, and MEF2D–CSF1R[7].
End of V4 Section
editv4:Clinical Significance (Diagnosis, Prognosis and Therapeutic Implications).Please incorporate this section into the relevant tables found in:
- Chromosomal Rearrangements (Gene Fusions)
- Individual Region Genomic Gain/Loss/LOH
- Characteristic Chromosomal Patterns
- Gene Mutations (SNV/INDEL)
- Diagnosis: Definitive diagnosis is based on two major gene expression signatures (DCOG/Erasmus MC and COG/St. Jude).
- DCOG/Erasmus MC incorporates hierarchal clustering of microarrays using a 110-gene probe set; this genetic signature frequently detected deletions in IKZF1, dic(9;20), and iAMP21 in BCR-ABL1-like B-ALL[8].
- COG/St. Jude employs predictive analysis of microarrays using a 257-gene probe set; this genetic signature demonstrated primarily activating kinase or cytokine receptor signaling alterations, in addition to IKZF1 deletions[9].
- Prognosis: In both pediatric and adult populations, BCR-ABL1-like B-ALL is associated with high rates of relapse and poor prognosis.
- The median 5-year overall survival rates for children with BCR-ABL1-like B-ALL, adolescents, and young adults was 72.8%, 65.8%, and 25.8%, respectively[10].
- Median 5-year-overall survival in adults was 22%, versus 64% in comparable patients with non-BCR-ABL1, non-BCR-ABL1-like, and non-MLL translocation B-ALL[11].
- Therapeutic Implications: Due to the aggressive nature of the disease, patients are often treated with high-intensity chemotherapy regimens, such as hyper-CVAD or an augmented Berlin-Frankfurt-Münster regimen[12].
End of V4 Section
Individual Region Genomic Gain/Loss/LOH
Put your text here and fill in the table (Instructions: Includes aberrations not involving gene rearrangements. Details on clinical significance such as prognosis and other important information can be provided in the notes section. Can refer to CGC workgroup tables as linked on the homepage if applicable. Please include references throughout the table. Do not delete the table.)
| Chr # | Gain, Loss, Amp, LOH | Minimal Region Cytoband and/or Genomic Coordinates [Genome Build; Size] | Relevant Gene(s) | Diagnostic, Prognostic, and Therapeutic Significance - D, P, T | Established Clinical Significance Per Guidelines - Yes or No (Source) | Clinical Relevance Details/Other Notes |
|---|---|---|---|---|---|---|
| EXAMPLE:
7 |
EXAMPLE: Loss | EXAMPLE:
chr7 |
EXAMPLE:
Unknown |
EXAMPLE: D, P | EXAMPLE: No | EXAMPLE:
Presence of monosomy 7 (or 7q deletion) is sufficient for a diagnosis of AML with MDS-related changes when there is ≥20% blasts and no prior therapy (add reference). Monosomy 7/7q deletion is associated with a poor prognosis in AML (add references). |
| EXAMPLE:
8 |
EXAMPLE: Gain | EXAMPLE:
chr8 |
EXAMPLE:
Unknown |
EXAMPLE: D, P | EXAMPLE:
Common recurrent secondary finding for t(8;21) (add references). | |
| EXAMPLE:
17 |
EXAMPLE: Amp | EXAMPLE:
17q12; chr17:39,700,064-39,728,658 [hg38; 28.6 kb] |
EXAMPLE:
ERBB2 |
EXAMPLE: D, P, T | EXAMPLE:
Amplification of ERBB2 is associated with HER2 overexpression in HER2 positive breast cancer (add references). Add criteria for how amplification is defined. | |
editv4:Genomic Gain/Loss/LOHThe content below was from the old template. Please incorporate above.
Monoallelic (often partial) deletion of the IKAROS transcription factor, encoded by IKZF1, is one of the most frequently observed genetic abnormalities in BCR-ABL1-like B-ALL, although this finding is not specific and not included in the definition[15].
End of V4 Section
Characteristic Chromosomal or Other Global Mutational Patterns
Put your text here and fill in the table (Instructions: Included in this category are alterations such as hyperdiploid; gain of odd number chromosomes including typically chromosome 1, 3, 5, 7, 11, and 17; co-deletion of 1p and 19q; complex karyotypes without characteristic genetic findings; chromothripsis; microsatellite instability; homologous recombination deficiency; mutational signature pattern; etc. Details on clinical significance such as prognosis and other important information can be provided in the notes section. Please include references throughout the table. Do not delete the table.)
| Chromosomal Pattern | Molecular Pathogenesis | Prevalence -
Common >20%, Recurrent 5-20% or Rare <5% (Disease) |
Diagnostic, Prognostic, and Therapeutic Significance - D, P, T | Established Clinical Significance Per Guidelines - Yes or No (Source) | Clinical Relevance Details/Other Notes |
|---|---|---|---|---|---|
| EXAMPLE:
Co-deletion of 1p and 18q |
EXAMPLE: See chromosomal rearrangements table as this pattern is due to an unbalanced derivative translocation associated with oligodendroglioma (add reference). | EXAMPLE: Common (Oligodendroglioma) | EXAMPLE: D, P | ||
| EXAMPLE:
Microsatellite instability - hypermutated |
EXAMPLE: Common (Endometrial carcinoma) | EXAMPLE: P, T | |||
editv4:Characteristic Chromosomal Aberrations / PatternsThe content below was from the old template. Please incorporate above.
Approximately half of cases demonstrate rearrangements resulting in overexpression of CRLF2[16]. These rearrangements are the result of either translocation of immunoglobin heavy chain enhance locus into CRLF2 (IGH-CRLF2—more commonly seen in adults) or through a cryptic deletion on chromosome X/Y involving the PAR1 psuedoautosomal region, resulting in fusion of CRLF2 to P2RY8 (more commonly seen in children). Very rare alternative translocations involving CRLF2 have also been observed.
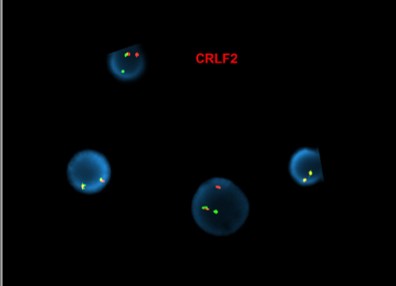
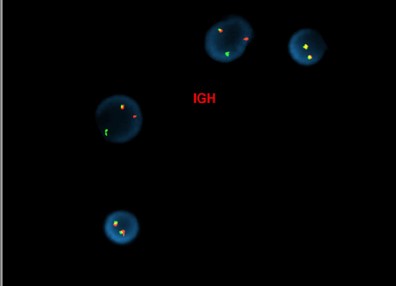
[Abnormal FISH results in interphase nuclei from a bone marrow sample using the CRLF2 dual-color, break-apart (Cytocell) and IGH dual-color, break-apart probes, reflective of IGH-CRLF2 fusion]

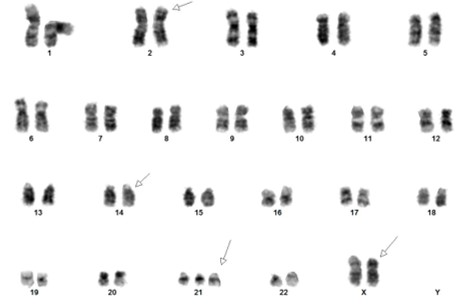
[Concurrent abnormal karyotype with trisomy 21 and a translocation involving chromosomes X, 14, and 2 in 9 of 13 cells available for analysis. Metaphase FISH with the IGH break-apart probe (Vysis) confirms the presence of 5’ IGH (green signal) on the abnormal chromosome Xp33.1 (CRLF2 locus), highly suggestive on an IGH-CRLF2 fusion rearrangement.
47,XX,+21c[4]/47,idem,der(X)t(X;14)(p33.1;q32),der(2)t(2;14)(p11.2;q11.2)t(X;14),der(14)t(2;14)[5]/46,XX[4].ish der(X)(5’IGH+),der(2)(3’IGH+)]
(Images courtesy of Fabiola Quintero-Rivera, MD)
End of V4 Section
Gene Mutations (SNV/INDEL)
Put your text here and fill in the table (Instructions: This table is not meant to be an exhaustive list; please include only genes/alterations that are recurrent or common as well either disease defining and/or clinically significant. If a gene has multiple mechanisms depending on the type or site of the alteration, add multiple entries in the table. For clinical significance, denote associations with FDA-approved therapy (not an extensive list of applicable drugs) and NCCN or other national guidelines if applicable; Can also refer to CGC workgroup tables as linked on the homepage if applicable as well as any high impact papers or reviews of gene mutations in this entity. Details on clinical significance such as prognosis and other important information such as concomitant and mutually exclusive mutations can be provided in the notes section. Please include references throughout the table. Do not delete the table.)
| Gene | Genetic Alteration | Tumor Suppressor Gene, Oncogene, Other | Prevalence -
Common >20%, Recurrent 5-20% or Rare <5% (Disease) |
Diagnostic, Prognostic, and Therapeutic Significance - D, P, T | Established Clinical Significance Per Guidelines - Yes or No (Source) | Clinical Relevance Details/Other Notes |
|---|---|---|---|---|---|---|
| EXAMPLE:EGFR
|
EXAMPLE: Exon 18-21 activating mutations | EXAMPLE: Oncogene | EXAMPLE: Common (lung cancer) | EXAMPLE: T | EXAMPLE: Yes (NCCN) | EXAMPLE: Exons 18, 19, and 21 mutations are targetable for therapy. Exon 20 T790M variants cause resistance to first generation TKI therapy and are targetable by second and third generation TKIs (add references). |
| EXAMPLE: TP53; Variable LOF mutations
|
EXAMPLE: Variable LOF mutations | EXAMPLE: Tumor Supressor Gene | EXAMPLE: Common (breast cancer) | EXAMPLE: P | EXAMPLE: >90% are somatic; rare germline alterations associated with Li-Fraumeni syndrome (add reference). Denotes a poor prognosis in breast cancer. | |
| EXAMPLE: BRAF; Activating mutations | EXAMPLE: Activating mutations | EXAMPLE: Oncogene | EXAMPLE: Common (melanoma) | EXAMPLE: T | ||
Note: A more extensive list of mutations can be found in cBioportal, COSMIC, and/or other databases. When applicable, gene-specific pages within the CCGA site directly link to pertinent external content.
editv4:Gene Mutations (SNV/INDEL)The content below was from the old template. Please incorporate above.
In addition to gene translocations, gain-of-function mutations in CRLF2 itself or in its partner gene, IL7RA, have been seen[17]. Alternative alterations activating kinase signaling occur, including activating mutations of FLT3, as well as focal deletions of SH2B3 (also known as LNK)[18].
Herold et al. in 2017 reported a wide variety of molecular alterations in BCR-ABL1-like B-ALL, which was shown to have statistically significant associations with alterations of IKZF1, CRLF2, JAK2, BTG1, and high CRLF2 expression[11].
End of V4 Section
Epigenomic Alterations
Not applicable
Genes and Main Pathways Involved
Put your text here and fill in the table (Instructions: Please include references throughout the table. Do not delete the table.)
| Gene; Genetic Alteration | Pathway | Pathophysiologic Outcome |
|---|---|---|
| EXAMPLE: BRAF and MAP2K1; Activating mutations | EXAMPLE: MAPK signaling | EXAMPLE: Increased cell growth and proliferation |
| EXAMPLE: CDKN2A; Inactivating mutations | EXAMPLE: Cell cycle regulation | EXAMPLE: Unregulated cell division |
| EXAMPLE: KMT2C and ARID1A; Inactivating mutations | EXAMPLE: Histone modification, chromatin remodeling | EXAMPLE: Abnormal gene expression program |
editv4:Genes and Main Pathways InvolvedThe content below was from the old template. Please incorporate above.
- IKAROS transcription factor: Deletion of IKZF1 results in activation of EBF1, MSH2, and MCL1, leading to B-cell leukemogenesis[19].
- CRLF2 overexpression: CRFL2 and its cofactor IL7RA form a receptor for thymic stromal-derived lymphopoietin that activates the JAK2-signal transducer and upregulates the transcription 5 pathway[17].
- Dysregulation of several tyrosine kinase signaling pathways (involving ABL1, ABL2, PDGFRB, CSF1, etc.) results in B-cell progenitor proliferation.
End of V4 Section
Genetic Diagnostic Testing Methods
- Flow cytometry for CRLF2 has been shown in some studies to be 100% concordant with FISH results[16].
- Next-generation sequencing is helpful for detecting copy number changes, single nucleotide variants, and gene fusions involving CRLF2, ABL1, ABL2, JAK2, etc.
- Gene expression profile algorithms, incorporating prediction analysis or hierarchical clustering of microarrays, provide the definitive diagnosis of BCR-ABL1-like B-ALL.
Familial Forms
Families with certain inherited variants of GATA3 (often seen in Native-American populations) are at increased risk of BCR-ABL1-like B-ALL[20].
Additional Information
Put your text here
Links
Pre-B ALL B-lymphoblastic leukemia/lymphoma with BCR-ABL1-like/Ph-like in Pathology Outlines (http://www.pathologyoutlines.com/topic/leukemiaprebbcrabl1like.html)
References
(use the "Cite" icon at the top of the page) (Instructions: Add each reference into the text above by clicking where you want to insert the reference, selecting the “Cite” icon at the top of the wiki page, and using the “Automatic” tab option to search by PMID to select the reference to insert. If a PMID is not available, such as for a book, please use the “Cite” icon, select “Manual” and then “Basic Form”, and include the entire reference. To insert the same reference again later in the page, select the “Cite” icon and “Re-use” to find the reference; DO NOT insert the same reference twice using the “Automatic” tab as it will be treated as two separate references. The reference list in this section will be automatically generated and sorted.)
- ↑ Akkari, Yassmine M. N.; et al. (2020-05). "Evidence-based review of genomic aberrations in B-lymphoblastic leukemia/lymphoma: Report from the cancer genomics consortium working group for lymphoblastic leukemia". Cancer Genetics. 243: 52–72. doi:10.1016/j.cancergen.2020.03.001. ISSN 2210-7762. PMID 32302940 Check
|pmid=value (help). Check date values in:|date=(help) - ↑ 2.0 2.1 2.2 Reshmi, Shalini C.; et al. (2017-06-22). "Targetable kinase gene fusions in high-risk B-ALL: a study from the Children's Oncology Group". Blood. 129 (25): 3352–3361. doi:10.1182/blood-2016-12-758979. ISSN 1528-0020. PMC 5482101. PMID 28408464.
- ↑ Zaliova, Marketa; et al. (2016-09). "Characterization of leukemias with ETV6-ABL1 fusion". Haematologica. 101 (9): 1082–1093. doi:10.3324/haematol.2016.144345. ISSN 1592-8721. PMC 5060025. PMID 27229714. Check date values in:
|date=(help) - ↑ Ernst, Thomas; et al. (2011-04). "Identification of FOXP1 and SNX2 as novel ABL1 fusion partners in acute lymphoblastic leukaemia". British Journal of Haematology. 153 (1): 43–46. doi:10.1111/j.1365-2141.2010.08457.x. ISSN 1365-2141. PMID 21391972. Check date values in:
|date=(help) - ↑ Duployez, Nicolas; et al. (2016-04). "NUP214-ABL1 fusion defines a rare subtype of B-cell precursor acute lymphoblastic leukemia that could benefit from tyrosine kinase inhibitors". Haematologica. 101 (4): e133–134. doi:10.3324/haematol.2015.136499. ISSN 1592-8721. PMC 5004396. PMID 26681761. Check date values in:
|date=(help) - ↑ Roberts, Kathryn G.; et al. (2014-09-11). "Targetable kinase-activating lesions in Ph-like acute lymphoblastic leukemia". The New England Journal of Medicine. 371 (11): 1005–1015. doi:10.1056/NEJMoa1403088. ISSN 1533-4406. PMC 4191900. PMID 25207766.
- ↑ Heim S & Mitelman F. Cancer Cytogenetics: Chromosomal and Molecular Genetic Aberrations of Tumor Cells. John Wiley & Sons, Incorporated: Chichester, United Kingdom. 2015.
- ↑ Den Boer, Monique L.; et al. (2009). "A subtype of childhood acute lymphoblastic leukaemia with poor treatment outcome: a genome-wide classification study". The Lancet. Oncology. 10 (2): 125–134. doi:10.1016/S1470-2045(08)70339-5. ISSN 1474-5488. PMC 2707020. PMID 19138562.
- ↑ Mullighan, Charles G.; et al. (2009). "Deletion of IKZF1 and prognosis in acute lymphoblastic leukemia". The New England Journal of Medicine. 360 (5): 470–480. doi:10.1056/NEJMoa0808253. ISSN 1533-4406. PMC 2674612. PMID 19129520.
- ↑ 10.0 10.1 Roberts, Kathryn G.; et al. (2014). "Targetable kinase-activating lesions in Ph-like acute lymphoblastic leukemia". The New England Journal of Medicine. 371 (11): 1005–1015. doi:10.1056/NEJMoa1403088. ISSN 1533-4406. PMC 4191900. PMID 25207766.
- ↑ 11.0 11.1 Herold, Tobias; et al. (2017). "Adults with Philadelphia chromosome-like acute lymphoblastic leukemia frequently have IGH-CRLF2 and JAK2 mutations, persistence of minimal residual disease and poor prognosis". Haematologica. 102 (1): 130–138. doi:10.3324/haematol.2015.136366. ISSN 1592-8721. PMC 5210243. PMID 27561722.
- ↑ Jain, Nitin; et al. (2017). "Ph-like acute lymphoblastic leukemia: a high-risk subtype in adults". Blood. 129 (5): 572–581. doi:10.1182/blood-2016-07-726588. ISSN 1528-0020. PMC 5290985. PMID 27919910.
- ↑ Tasian, Sarah K.; et al. (2012). "Aberrant STAT5 and PI3K/mTOR pathway signaling occurs in human CRLF2-rearranged B-precursor acute lymphoblastic leukemia". Blood. 120 (4): 833–842. doi:10.1182/blood-2011-12-389932. ISSN 1528-0020. PMC 3412346. PMID 22685175.
- ↑ Iacobucci, Ilaria; et al. (2016). "Truncating Erythropoietin Receptor Rearrangements in Acute Lymphoblastic Leukemia". Cancer Cell. 29 (2): 186–200. doi:10.1016/j.ccell.2015.12.013. ISSN 1878-3686. PMC 4750652. PMID 26859458.
- ↑ Boer, Judith M.; et al. (2015). "BCR-ABL1-like cases in pediatric acute lymphoblastic leukemia: a comparison between DCOG/Erasmus MC and COG/St. Jude signatures". Haematologica. 100 (9): e354–357. doi:10.3324/haematol.2015.124941. ISSN 1592-8721. PMC 4800707. PMID 26045294.
- ↑ 16.0 16.1 Konoplev, Sergej; et al. (2017). "CRLF2-Positive B-Cell Acute Lymphoblastic Leukemia in Adult Patients: A Single-Institution Experience". American Journal of Clinical Pathology. 147 (4): 357–363. doi:10.1093/ajcp/aqx005. ISSN 1943-7722. PMID 28340183.
- ↑ 17.0 17.1 Quesada A, Reynolds M, Jorgensen JL, et al. Cytokine receptor-like factor 2 (CRLF2) expression in precursor B-lymphoblastic leukemia. International Clinical Cytometry Society e-Newsletter. 2014;5(1).
- ↑ Tosi S & Reid AG. The Genetic Basis of Haematological Cancers. John Wiley & Sons, Incorporated: Chichester, United Kingdom: 2016.
- ↑ van der Veer, Arian; et al. (2013). "Independent prognostic value of BCR-ABL1-like signature and IKZF1 deletion, but not high CRLF2 expression, in children with B-cell precursor ALL". Blood. 122 (15): 2622–2629. doi:10.1182/blood-2012-10-462358. ISSN 1528-0020. PMC 3795461. PMID 23974192.
- ↑ Perez-Andreu, Virginia; et al. (2013). "Inherited GATA3 variants are associated with Ph-like childhood acute lymphoblastic leukemia and risk of relapse". Nature Genetics. 45 (12): 1494–1498. doi:10.1038/ng.2803. ISSN 1546-1718. PMC 4039076. PMID 24141364.
Notes
*Primary authors will typically be those that initially create and complete the content of a page. If a subsequent user modifies the content and feels the effort put forth is of high enough significance to warrant listing in the authorship section, please contact the Associate Editor or other CCGA representative. When pages have a major update, the new author will be acknowledged at the beginning of the page, and those who contributed previously will be acknowledged below as a prior author.
Prior Author(s):
*Citation of this Page: “B-lymphoblastic leukaemia/lymphoma with BCR::ABL1-like features”. Compendium of Cancer Genome Aberrations (CCGA), Cancer Genomics Consortium (CGC), updated 10/15/2025, https://ccga.io/index.php/HAEM5:B-lymphoblastic_leukaemia/lymphoma_with_BCR::ABL1-like_features.